- Open access
- Published: 11 March 2016

Study of Leishmania pathogenesis in mice: experimental considerations
- Corinne Loeuillet 1 ,
- Anne-Laure Bañuls 1 &
- Mallorie Hide 1
Parasites & Vectors volume 9 , Article number: 144 ( 2016 ) Cite this article
7882 Accesses
89 Citations
1 Altmetric
Metrics details
Although leishmaniases are endemic in 98 countries, they are still considered neglected tropical diseases. Leishmaniases are characterized by the emergence of new virulent and asymptomatic strains of Leishmania spp. and, as a consequence, by a very diverse clinical spectrum. To fight more efficiently these parasites, the mechanisms of host defense and of parasite virulence need to be thoroughly investigated. To this aim, animal models are widely used. However, the results obtained with these models are influenced by several experimental parameters, such as the mouse genetic background, parasite genotype, inoculation route/infection site, parasite dose and phlebotome saliva. In this review, we propose an update on their influence in the two main clinical forms of the disease: cutaneous and visceral leishmaniases.
Leishmaniases are caused by pathogens of the genus Leishmania and are characterized by an important clinical and epidemiological diversity. According to the World Health Organization (WHO), leishmaniases occur in four continents and are endemic in 98 countries [ 1 ]. The causative parasites are classified according to genetic, biological and immunological criteria. More than 30 Leishmania spp. are known of which 20 are pathogenic for humans (for a taxonomic table, see [ 2 ]). In humans, this disease can have several clinical manifestations of variable severity: mucosal leishmaniasis (a mutilating disease), diffuse cutaneous leishmaniasis (a long-lasting disease due to a deficient cell-mediated immune response), cutaneous leishmaniasis (disabling with multiple lesions) or visceral leishmaniasis, which is fatal if untreated (see for reviews [ 3 , 4 ]). Leishmaniasis can also be asymptomatic in humans [ 5 , 6 ] and in animal reservoirs [ 7 , 8 ]. It is worth noting that a single species of Leishmania may cause several disease forms [ 3 ].
Drug-resistant Leishmania isolates or emerging virulent strains are often described [ 9 , 10 ]. It is crucial to understand how these new strains disseminate and are controlled by the host immune system to fight against them. To investigate the pathogenic diversity, the impact of the host genetic background and of the Leishmania genotypes, animal models are widely used. Classically, in infected animals, parasite-activated CD4 + T cells rapidly proliferate in the lymph nodes, differentiate and secrete specific cytokines. Th1 cells secrete IL2, IFNγ and TNFα, leading to macrophage activation and parasite elimination. On the other hand, the Th2 response is associated with IL4, IL5 and IL13 production and with parasite proliferation (for review see [ 11 ]). When studying a newly isolated strain, the experimental settings have to be carefully designed and several parameters must be taken into account. The objective of this review is to summarize results on the pathogenic mechanisms in mice infected by Leishmania spp. We will focus on the two main clinical forms: visceral leishmaniasis (VL) and cutaneous leishmaniasis (CL). We will first describe the experimental data on the influence of the genetic background in mouse models of VL and CL caused by Leishmania donovani and L. infantum and of CL caused by L. major , L. mexicana and L. tropica . Then, data obtained in mouse models of VL by L. infantum and of CL by L. major will be reviewed, particularly: (i) the immune cells involved and the associated-immune response and (ii) the parameters (mouse and parasite genotypes, parasite dose and inoculation route) that influence the infection outcome.
Visceral leishmaniasis
Human VL is the most severe form of this disease and occurs when infected people are left untreated. Ninety percent of reported cases of VL are found in Bangladesh, Brazil, India, Nepal and Sudan (WHO). Approximately, 0.2 to 0.4 million cases of VL are reported each year and VL causes 20,000 to 40,000 deaths annually [ 1 ]. It is caused by parasites of the Leishmania donovani complex: L. donovani , L. archibaldi and L. infantum (syn. L. chagasi ). The main causative agents of VL in humans are L. donovani and L. infantum , whereas L. infantum can cause CL (see below) and is the main VL-causing parasite in dogs, which are considered to be Leishmania spp. reservoirs [ 3 ]. Hereafter, we will describe the main experimental parameters that need to be taken into account when studying visceralizing Leishmania strains.
Mouse genetic background
Susceptibility and resistance to Leishmania infection in humans and in mice are regulated by genetic determinants [ 12 ]. In the mouse, two main loci have been described: Lsh and H 2 (the major histocompatibility complex). The Lsh locus on mouse chromosome 1, with Nramp1 ( Slc11a1 ) as the main candidate gene [ 13 ], influences the natural resistance to L. donovani infection (control of parasite load in liver and spleen) [ 14 ]. In mice harboring wild type Scl11a1 , such as the CBA mouse strain, parasite proliferation in liver is hindered. Conversely, in mice with mutant Scl11a1 , such as the BALB/c and C57BL/6 strains, parasite growth is unrestrained [ 15 ]. The H 2 locus [ 16 ] is involved in the development of the adaptive immunity and overcomes the innate susceptibility to L. donovani caused by Scl11a1 mutations [ 15 ]. Like for L. donovani , L. infantum primary infection is also initially controlled by the Lsh locus and then by the H 2 locus. Both loci are involved in the development of the acquired immune response [ 17 ]. For instance, L. infantum susceptible mice (with mutant Scl11a1 ) harboring the H2-b or H2-r alleles, show a “cure” phenotype associated with a strong reduction of the parasitic load in liver. Conversely, H2-d, H2-q and H2-f susceptible mice are characterized by a “non-cure” phenotype. Mice can thus be classified in four phenotypic categories: resistant, susceptible and when susceptible, in cure and non-cure (Table 1 ). Non-cure mice, such as the BALB/c strain, will spontaneously progress to a chronic phase of the disease without total parasite clearing. Conversely, in cure mice, such as the NMR1 strain, the parasite load is very low and parasites can be fully eliminated in some cases.
In conclusion, it is clear that the mouse genetic background influences the Leishmania infection outcome and this feature must be taken into account when designing experiments. Although BALB/c mice will not die of VL (differently from untreated humans), they can be used to study the immunopathology changes occurring during VL. Indeed, they present clinical features of human VL, such as hepatosplenomegaly or disruption of splenic tissue. However, the infection outcome depends also on the ability of the infected mice to induce a good Th1 immune response as well as on the inoculation route and injected dose, as discussed hereafter.
Immune control of infection
From an immunological point of view, the main feature of VL is its organ specificity. Indeed, studies on VL progression in mice highlighted that the major responding tissues (spleen and liver) present distinct patterns in term of immune response and parasite control. When injected intravenously in BALB/c mice, more than 95 % of L. infantum promastigotes are rapidly cleared from the circulation by highly phagocytic macrophages of the spleen marginal zone [ 18 ]. After pathogen uptake, macrophages exert their leishmanicidal activity through nitric oxide (NO) synthesis. During the course of infection, the parasite burden is progressively controlled [ 19 ] with a peak of parasitemia at week 8 of infection [ 20 ]. In the spleen, the course of parasite burden reflects the cytokine production kinetics and the activation of the different classes of immune cells. At the beginning of infection (first 4 weeks), parasite replication is associated with the immune cell inability to produce IFNγ and IL2 (macrophage-activating cytokines), whereas production of IL4 or IL5 is conserved [ 21 , 22 ]. Conversely, IL10 and TGFβ (macrophage-inhibitory cytokines) are produced by cells of the spleen marginal zone and of the red pulp, respectively. IL10 and TGFβ are thought to contribute to the establishment of infection and parasite replication [ 19 ]. After the first 4 weeks of infection, CD4 + T, CD8 + T and natural killer (NK) cells recover their capacity to produce IFNγ, thus promoting the macrophage microbicidal activity with NO synthesis and control of granuloma formation in liver (see next paragraph) and ultimately parasite burden reduction [ 21 ]. In synergy with IFNγ, IL17A also contributes to macrophage activation with NO production, leading to parasite clearance [ 23 ]. Nevertheless, infection in the spleen is maintained during the entire VL course. This parasite persistence may be due to sustained TGFβ production by CD4 + CD25 + T cells (Treg) that contributes to immunosuppression [ 24 ].
In liver, the infection time course is different. After injection in the lateral tail vein, promastigotes invade the resident macrophages, Kupffer cells and dendritic cells where they become amastigotes and replicate. During acute infection (first 2 weeks), parasite growth is uncontrolled, due to elevated TGFβ levels and ineffective Th1 response [ 25 ]. This correlates with the peak of parasite burden observed between 2 and 8 weeks post-infection. Liver infection is then resolved through the formation of granulomas that are characterized by parasitized Kupffer cells surrounded by a mantle of lymphocytes [ 26 ]. Finally, after 8 weeks of infection, amastigotes are almost absent in granulomas and infection is resolved [ 26 ].
In VL, the organ-specific immunity with elimination of parasites in liver and their persistence in the spleen directly reflects the observed immune response. This chronic status is critical because the host becomes more susceptible to secondary infections. Thus, to develop and evaluate new vaccines or therapies against leishmaniases, it is crucial to determine the role of each immune cell type in the establishment of the cellular immune interplay resulting in the control of the infection.
For VL, a thorough understanding of the mechanisms underlying the protective immune response in liver and the failure in spleen would allow the development of new candidate vaccines and of new strategies or treatments to eliminate the parasite in infected people.
Tissue of origin and parasite genotype
Another question is whether the disease pattern differs depending on the tissue of origin of the parasites used to infect mice (isolated from cutaneous or visceral forms in humans). Comparison of the infection profiles (parasite burden in spleen and liver) in BALB/c mice of 22 L. infantum strains isolated from patients with CL or VL revealed a great variability of infection profiles (progressive, controlled or undetectable infection) [ 27 ]. The parasite zymodeme and the host immune status did not have any influence on the infection profile. However, no strain of cutaneous origin caused a visceralizing form in mice [ 27 ]. In susceptible mice, histopathological analysis of the liver revealed no difference whatever the L. infantum strain used, although a more pronounced liver granulomatous response was observed with visceralizing strains [ 28 ]. Interestingly, the visceralizing and infection profiles were preserved in BALB/c and C57BL/6 mice, both susceptible to infection with a non-cure and a cure profile respectively, suggesting a clear role of parasite factors on the infection outcome. This was confirmed by the finding that the infection kinetics were similar following experimental infection of immunodeficient C.B-17 SCID and congenic BALB/c mice, although the parasite load was higher in immunodeficient animals [ 29 ].
Moreover, L. infantum strains belonging to the same zymodeme (MON-1) can lead to different infection profiles in mice. The most pathogenic strains induced splenomegalia and higher parasite load in spleen and liver associated with higher IgG1, TGFβ and reduced IFNγ production [ 30 ]. This effect was not related to the host species (dog or human) from which the parasites were isolated [ 30 ]. These observations highlight the intra-strain specific virulence variability and confirm that in VL, parasite virulence is a clonal or inbred dominant trait within each strain (infection study of 11 clones derived from three strains of known virulence) [ 31 ].
The influence of the host immune status on the strain virulence is not well known. Indeed, strains isolated from immunosuppressed patients display either high [ 27 ] or low virulence [ 30 ].
In conclusion, it is difficult to extend experimental results in mouse models obtained with a given Leishmania strain to other strains even if they are genetically related or responsible for the same clinical form. Therefore, it is crucial to analyze all new parasite isolates and the corresponding immune response in mice. Such studies are particularly recommended for parasite stocks that cause large epidemics or non-pathogenic and silent leishmaniasis forms in humans. Prospective studies in endemic areas also are important to better understand the basis of the protective immune response in humans and to develop specific and more efficient treatments.
Inoculation route and parasite dose
Several inoculation routes have been tested (Table 2 ). It appears that the parasite load (reflecting the immune response strength) depends not only on the parasite dose that is inoculated, but also on the chosen injection route.
For the LIVT-1 strain, the subcutaneous route seems to be less efficient (based on the parasite load in the spleen) than the intravenous one, when mice are inoculated with medium doses of parasites (10 5 ); however, it has been not demonstrated for higher doses (10 6 and 10 7 ) [ 32 ]. Moreover, the parasite load in the liver is higher following intravenous inoculation compared to the subcutaneous route and the response is dose-dependent [ 32 ].
A clear dose effect on parasite load was observed in BALB/c mice inoculated subcutaneously with L. infantum [ 33 ]. Mice inoculated with low doses (10 3 ) showed a minimal infection associated with a Th1 response (IFNγ and NO production in spleen), whereas high doses (10 7 ) led to high parasite burden in spleen and lymph nodes as well as to a Th2 response [ 33 ].
By assessing several inoculation routes, Kaur et al. showed that the subcutaneous route is less efficient than the intradermal, intraperitoneal and intracardiac routes in term of liver parasite load [ 34 ]. Again, a clear dose effect on parasitemia was detected. High doses inoculated by intracardiac injection led to the highest liver parasite load and were associated with a Th2 immune response [ 34 ]. However, the Th1 immune response necessary for the establishment of resistance in BALB/c mice was strongest when mice were inoculated with low doses of parasites via the subcutaneous or intradermal routes. This was associated with maximum IFNγ production and high level of IgG2a [ 33 , 34 ]. It is worth noting some discrepancies in studies assessing the same dose and the same route. For example, differently from Rosypal et al., Oliviera and colleagues demonstrated that, at high dose (10 7 ), parasite load is higher when mice are inoculated subcutaneously rather than intravenously. Overall, it is difficult to compare different studies because of the different parasite strains and parasite load determination methods. This stresses again that results obtained with a given stock cannot be fully transposed to another one. Moreover, although all the studies summarized here were performed using BALB/c mice, we cannot assume that the animals were genetically identical and this could account for the different responses sometimes observed after Leishmania infection.
The intracardiac route is responsible for the development of Th2 immune response that is characterized by IL4 production and increased levels of IgG1 and is associated with IL10 production by Treg cells, thus allowing the establishment of a persistent infection [ 24 , 34 ]. Although another study described a quite different immune response after intracardiac infection [mixed activating (IFNγ and TNFα)/deactivating (TGFβ) cytokine response] [ 35 ], this route seems to be effective in term of infection persistence. Indeed, intracardiac inoculation leads to progressive VL with parasite persistence in the spleen 4 months after the infection and accompanied by massive splenomegaly [ 35 ].
Intravenous inoculation seems to lead to effective infection (based on parasite load) whatever the dose used, with parasite persistence in spleen and liver when high doses are injected [ 36 ]. Liver lesions are prominent in intravenously inoculated mice, whereas they are almost inexistent in mice injected subcutaneously. However, heterogeneity is observed among animals as already described for spleen parasite load [ 37 ]. Liver granuloma formation seems to depend on the dose used: high numbers of mature and sterile granulomas are observed when mice are inoculated with low doses, whereas no parasite clearance is detected after injection of high doses [ 36 ]. However, it is worth noting that only mice inoculated with low doses could mount a protective response in the liver and bone marrow, associated with parasite clearance.
Intradermal inoculation of high parasite doses (10 7 ) leads to chronic infection with parasite persistence in spleen and lymph nodes, Th1/Th2 cytokines production (IFNγ, IL4) and partial clearance in the liver [ 38 ]. When inoculated with low (10 3 ) or medium (10 5 ) doses, mice are effectively infected, but present a lower parasite load and a Th1-dominant immune response [ 33 ].
In conclusion, although the subcutaneous or intradermal routes mimic the natural infection routes, they require high doses of parasites to cause a good and persistent infection. Lower doses can be recommended for vaccination protocols because the generated immune response seems to be sufficient for long-term protection. One study reported that the intraperitoneal route leads to higher homogeneity of infection in term of parasite load and thus might be more appropriate to study new isolates [ 37 ]. Conversely, the intravenous route leads to heterogeneous parasite loads among animals and, consequently, errors in the interpretation of the results could occur when analyses are performed on pooled animals.
Influence of phlebotome saliva
Leishmania parasites are classically injected in the skin together with saliva by an infected sandfly during a blood meal. For VL, studies on phlebotome saliva are mainly focused on its modulatory effect on the immune response or as a vaccine candidate rather than on its influence on the primary infection outcome (for review see [ 39 ]). To our knowledge, only two studies reported that, differently from CL (see below), salivary extracts do not to have a significant role in VL establishment in dogs and mice [ 38 , 40 ]. Dogs intradermally inoculated with L. chagasi stationary promastigotes together with or without Lutzomyia longipalpis salivary gland extract did not show any infection, suggesting that the presence of salivary proteins is not sufficient for effective infection through the intradermal route [ 40 ]. In BALB/c mice, intradermal injection of L. infantum promastigotes with or without sandfly salivary extracts led to effective visceralization in both cases, showing that salivary products do not influence the infection course (parasite burden in spleen, liver and lymph nodes) [ 38 ]. Further studies are needed to validate these results and to determine the potential effect of sandfly salivary components on L. infantum visceralization capacity and infection outcome.
In conclusion, when studying VL in mice, it is important to consider the parasite dose as well as the route of inoculation because they clearly influence the development of the T helper response and consequently the infection outcome. The study of the chronic phase of infection requires an inoculation route (intravenous or intradermal) that allows the long-term establishment of the infection. Conversely, for studying the acute phase of infection, the inoculated dose needs to be precisely controlled because a more intense immune response is observed with high doses. For vaccination studies, low doses of parasites (10 4 ) can be used because the elicited immune response can protect on the long term.
Cutaneous leishmaniasis
The incidence of human cutaneous leishmaniasis (CL) is of about 0.7 to 1.2 million cases each year [ 1 ] (WHO, 2014). According to the WHO, 90 % of CL cases occur in Afghanistan, Brazil, Iran, Peru, Saudi Arabia and Syria. It is usually a self-healing disease, but in the presence of multiple lesions, CL can lead to lifelong aesthetic stigma [ 4 ]. CL is caused by several Leishmania species: L. major , L. tropica , L. mexicana , L. amazonensis , L. braziliensis and L. guyanensis [ 2 , 3 ]. L. infantum , a classically visceralizing species, can also present an unusual skin tropism, thus rarely causing limited cutaneous lesions at the biting site [ 41 ].
Mice genetic background
Studies on the infection phenotype in mice with different genetic background and in inbred congenic mice have allowed the identification of several loci involved in the infection control (Table 3 ). Analysis of the lesion size in the progeny from crosses between resistant and susceptible mice following intradermal infection with L. tropica led to the hypothesis that susceptibility (C57BL/6 x Balb/c) or resistance (C3H/HeN x P/J) to CL could be controlled by a unique locus with minor influences by other genes [ 42 , 43 ]. However, the finding that not all animals with cutaneous lesions will develop a systemic infection suggests that different genes may be involved in this two forms of disease [ 42 , 43 ].
High throughput genetic screening revealed a complex picture. Indeed, some loci that control CL are common to several Leishmania spp., while others are specific. The Lsh1 locus on chromosome 1 can control the outcome of cutaneous infections caused by L. mexicana , but not by L. major [ 44 , 45 ]. The H2 locus also influences the infection outcome, but more weakly than the Lsh1 locus. Depending on the HLA allele, mice have been classified as resistant (H2-r, -s, -a, -k) or susceptible (H2-b, -d, -q), with slowing resolving lesions following infection by L. major , L. tropica or L. mexicana [ 46 – 48 ].
Studies based on genome-wide analysis of resistance versus susceptibility phenotypes in the offspring of various intercrosses allowed the identification of different loci involved in mouse susceptibility or resistance to CL. For instance, the scl -1 and -2 (for susceptibility to cutaneous leishmaniasis) loci on chromosome 11 [ 49 ] are involved in the control of mouse susceptibility. The scl - 1 locus controls the healing versus non-healing responses to L. major and the scl-2 is responsible for the development of cutaneous lesions induced by L. mexicana [ 49 ]. On the other hand, others loci are involved in mouse resistance as the Lmr -1, -2 and -3 (for Leishmania major resistance) loci that contribute to control skin lesion healing (for review see: [ 50 , 51 ]. Currently, more than 30 loci have been identified as involved in the complex control of L. major -induced CL. Indeed, a single Quantitative Trait Locus (QTL) does not account for the overall phenotype variance, but rather is responsible for the control of a specific infection or pathogenic aspect. For example, Lmr -5 regulates parasite load in spleen and Lmr -20 in lymph nodes, whereas Lmr -5 and Lmr -21 control the development of skin lesions [ 52 ]. Thus, contrary to VL where only two main loci ( Lsh1 and H2 ) are involved, CL outcome is regulated by the combination of several loci. The susceptibility/resistance of different mouse strains to CL is summarized in Table 4 . The highly susceptible BALB/c mice and the resistant C57BL/6 J mice are widely used to study both the genetics and biology of the host response to CL.
Due to the complex control of CL pathogenesis, we decided to focus mainly on L. major because it is the most studied strain. In experimental settings for in vivo studies, some parameters, described hereafter, need to be considered.
Differently from VL induced by L. donovani complex species where a mixed Th1/Th2 response is observed during the infection course, the outcome of L. major induced-CL depends on the development of polarized Th1 or Th2 responses associated with resistance or susceptibility, respectively [ 53 ]. Indeed, in resistant mice (C57BL/6), a Th1-oriented immune response, associated with IFNγ, IL2 and IL12 production, is clearly observed. At the infection site, few parasites remain viable thanks to the presence of CD4 + CD25 + regulatory T cells that produce IL10 [ 54 ]. Conversely, susceptible mice (BALB/c) develop a Th2 immune response with IL4 production, leading to the development of uncontrolled lesions and disseminated visceral infection. Treg cells that produce IL4 and IL10 cytokines also play a role in disease promotion by expanding, or regulating the Th2 population [ 55 ]. In these mice, lesion severity is also associated with the production of IL17 that promotes neutrophil immigration and thus lesion progression [ 56 ].
In experimental mouse models of CL, several cell types are found in the lesions: neutrophils [ 57 ], macrophages [ 58 ], eosinophils [ 58 ], lymphocytes [ 59 ], mast cells [ 60 ] and NK cells [ 61 ]. These cells play a role during the different phases of the infection: (i) silent phase (no lesion formation or inflammation detection) with parasite invasion of skin resident macrophages and neutrophils; (ii) lesion development associated with migration and activation of cells of the innate immune system (mast cells, neutrophils, monocytes); (iii) lesion involution with migration of dendritic cells and T cells; and (iv) chronic phase characterized by lesion resolution and associated with parasite persistence, mainly in macrophages, and lifelong immunity [ 58 ].
Thus, immunity to L. major depends on multiple cell types that cooperate for the development of an effective and protective immune response. Understanding their respective role and how to modulate their function could lead to new therapeutic approaches for immunization and long-lasting protection. For example, as dendritic cell activation is required for protective immunity, vaccines using infected or antigen-loaded dendritic cells could lead to the development of a specific and efficient immune protective response.
The remaining part of this review will focus on the parameters that influence the mouse immune response to CL: parasite genotype, parasite dose, site of the intradermal inoculation (ear dermis, dorsal skin and hind footpad), and associated adjuvant (saliva).
Parasite genotype
Few reports have assessed the natural virulence variability of different L. major strains in the same mouse model and the associated immune response. Li et al. compared the infection outcome of two clones derived from the same L. major strain. They found that the avirulence of the S2 clone, characterized by spontaneous lesion healing, was not correlated with its capacity to infect macrophages or the inoculated dose, but with parasite factors [ 62 ]. The study of the infection outcome in BALB/c mice inoculated with 19 L. major strains (12 from Tunisia, zymodeme 25; and 7 from the Middle East, zymodeme 26, 68, 70 or 103) revealed a large heterogeneity of disease severity (footpad lesion size) [ 63 ]. Interestingly, all Middle East strains presented high or intermediate virulence, whereas most Tunisian strains (10/12) showed lower virulence. This was correlated with their pathogenicity in humans. Higher virulence could be associated with a greater capacity to infect bone marrow-derived macrophages, faster growth in culture and the induction of a stronger Th2 response in vivo . Moreover, the in vitro study of two of these L. major clones (zymodeme 25) highlighted their different capacity of human dendritic cell invasion, a feature that could modulate the innate immune response [ 64 ].
The heterogeneity of lesion size in function of the parasite genotype was confirmed by another study in BALB/c mice [ 65 ]. In addition, these authors observed reproducible differences in lymph node parasite burden, depending on the L. major strain, at week 8 after inoculation. Specifically, the highest pathogenicity (based on the parasite load) was associated with induction of the Th2 immune response, whereas strains with intermediate or low pathogenicity elicited predominantly a Th1 immune response. Recently, the study of four Iranian strains in BALB/c mice highlighted their high diversity of lymph node parasite burden and cytokine expression and confirmed that the strain causing the lowest parasite burden induced mainly a Th1 response [ 66 ].
These results clearly demonstrate the importance of the parasite genotype in CL development, although one study suggested that high parasite dose could be the only important determinant of the Th1/Th2 response, independently of the parasite or the mouse genotypes [ 67 ].
Parasite dose
Concerning the inoculated dose (Table 5 ), an initial study found that in susceptible BALB/c mice, no clinical sign was visible (such as increase in footpad size) following subcutaneous inoculation of low parasite doses (10 2 to 10 3 parasites). Conversely, inoculation of high doses (10 5 to 10 7 parasites) led to significant footpad enlargement [ 68 ]. This dose-dependent effect was confirmed in other studies. For example, lesions were apparent in all BALB/c mice injected with 10 4 (MHOM/IR/-/173 strain) or 10 6 (MHOM/IL/80/Friedlin strain) parasites [ 67 ]. Uzonna et al. confirmed that CL severity (asymptomatic with no lesion but IgG2a response > IgG1; apparent lesions with identical IgG1 and IgG2a levels; large lesions and even foot loss) in BALB/c mice depends on the injected dose [ 69 ]. Moreover, subclinically infected BALB/c mice are resistant to a secondary pathogenic infection (10 6 parasites, footpad injection) and, thus, could be used in vaccination strategies [ 69 ]. However, not all mice infected with low parasite doses will develop a subclinical form of CL. Indeed, susceptible BALB/c mice inoculated with 10 2 parasites showed significant pathology (antibody response and parasite detection in lymph nodes) and then progressed to a chronic phase where lesions stop increasing in size and eventually will resolve [ 69 , 70 ]. In C57BL/6 mice, whatever the dose (10 2 to 10 7 ), lesions resolved and this was associated with the induction of a Th1 immune response (tested for the 10 2 and 10 6 doses) [ 70 ]. However, in another study, the classical clinical response (lesion healing and Th1 immune response) was observed only when C57BL/6 mice were inoculated with high parasite dose (10 6 ). Conversely, in animals inoculated with low doses (10 3 ), an unexpected, but transient Th2 response occurred first and was then reversed by the activation of IFNγ − producing CD8 + T cells [ 71 ].
In summary, the inoculated dose can influence the immune response and thus CL severity, which is also dependent on the mouse genetic background. Specifically, in susceptible mice (i.e. BALB/c strain), high doses (10 5 to 10 7 parasites) lead to persistent infection associated with a Th2 immune response. Low doses (10 1 or 10 2 parasites) induce a Th1 response and thus could be used in vaccination studies. However, a “serodeconversion” may occur several (9 to 18) months after infection. With intermediate doses (10 3 -10 4 ), a mixed Th1-Th2 immune response is observed.
In resistant mice (i.e., C57BL/6 strain), effective lesion development is observed only with high doses (10 6 -10 7 ). Inoculation of low doses (100 metacyclic promastigotes) at a dermal site (for instance, ear dermis) is recommended to mimic the natural transmission and induces two distinct disease phases [ 58 , 72 ]. First, a clinically silent phase occurs during the first 4–6 weeks and is characterized by the absence of lesions and the increase of the parasite load. During the second phase, lesions develop (footpad swelling) associated with immune cell infiltration at the site of infection. Concomitantly, Th1 cells expand in the draining lymph nodes, ultimately leading to parasite burden reduction and lesion healing.
- Inoculation site
In experimental studies on CL, only the intradermal route of inoculation is used, but at different sites: hind footpad, ear pinna and tail base (Table 6 ). Depending on the inoculation site, the clinical signs (lesion size and immune response) clearly differ. Moreover, the cytokine production profile does not always explain the disease severity [ 73 , 74 ].
In BALB/c mice, severe, non-healing lesions were observed whatever the inoculation site [ 73 , 74 ], and they were associated with a classical Th2 immune response [ 73 , 74 ].
In SWR mice, inoculation at the base of the tail led to the development of large non-healing lesions, whereas self-healing lesions were observed following inoculation in the hind footpad [ 73 ]. Unexpectedly, inoculation at the tail base tail induced a Th1 immune response, which is normally associated with self-healing lesions.
This dichotomy was also observed in resistant mice, such as the C57BL/6J strain. After inoculation in the ear pinna, the classical Th1 response was associated with the development of small, self-healing lesions. Conversely, parasite inoculation at the tail base, induced a Th2 response that was unexpectedly associated with lesion healing [ 74 ].
C3H/HeN and DBA/2 mice were resistant to CL caused by ear pinna inoculation of Leishmania parasites, but presented intermediate disease (C3H) or were fully susceptible (DBA/2) when inoculated at the tail base [ 74 ]. Again, no clear correlation was found between the type of immune response and the cutaneous lesion severity. For instance, C3H/HeN mice were fully resistant to L. major infection through the ear pinna, but no Th1 response could be detected.
CBA/H mice were resistant to L. major infection through any inoculation site, with an immune response similar to controls [ 74 ].
Thus, besides the parasite dose and mouse genetic background, the infection site also affects CL severity.
Several studies assessed the role of the vector saliva in CL development.
When L. major parasites were inoculated with saliva from Lutzomyia longipalpis , the size of cutaneous lesions in CBA and BALB/c mice was five to ten times bigger and contained at least 5000 times more parasites than in controls (no saliva) [ 75 ]. Moreover, when inoculated at low doses, parasites survived only when co-injected with saliva [ 75 ]. This disease exacerbation effect was observed in other mouse strains (susceptible, intermediate susceptible or resistant) and was more pronounced in resistant CBA and C57BL/6 mice [ 76 ]. Salivary gland extracts from L. longipalpis exacerbated CL following infection by L. major [ 78 ] or by L. braziliensis [ 57 , 78 ]. It has been demonstrated that maxadilan, a salivary vasodilator, is responsible for the disease-exacerbation effect of saliva from L. longipalpis [ 79 ].
Phlebotomus papatasi saliva also can increase lesion size, but with a less pronounced effect than saliva from L. longipalpis [ 76 ]. Moreover, inoculation of parasites with P. papatasi saliva in resistant CBA mice led not only to bigger lesion size, but also to higher parasite burden in lesions that was associated with a modulation of the immune response (decrease of Th1 factors and increase of Th2-associated IL4 production) [ 80 ]. When mimicking natural infection by injection of low dose of parasites in the ear dermis, co-inoculation of P. papatasi saliva promoted lesion development (earlier and higher parasitemia) in BALB/c and also in C57BL/6 mice [ 81 ]. This was associated with Th2 immune response induction and IL4 production.
Additional experimental parameters
To our knowledge, only one study in BALB/c mice described the influence of L. major infectious stage on (i) lesion development and ulceration and (ii) on the type of immune response [ 82 ]. Specifically, after inoculation of high doses (10 6 ) of metacyclic promastigotes (infective stage) in the ear dermis (pinna), lesions were detectable 1 week after infection and became ulcerate after 4 weeks. Following inoculation of high doses (10 6 ) of log-phase parasites (division stage), detectable lesions and ulcerations were observed after three and 10 weeks, respectively. Moreover, inoculation of log-phase parasites led to better activation of lymph node CD4 + T cells (IFNγ production) than inoculation of metacyclic promastigotes, at least during the early stages of infection (16 h and 3 days post-inoculation).
In addition, the presence of apoptotic promastigotes in the infectious inoculum is important for the parasite intracellular survival and, thus, for disease development in vivo [ 83 ]. Although this parameter must be further investigated, it has to be taken into account when designing in vivo experiments.
In summary, when studying CL, the route of infection, the infectious parasite stage, the inoculated dose or the adjuvant, the mouse resistance or susceptibility to infection are all essential parameters to be taken into account because they can substantially influence the issue of in vivo experiments.
Conclusions
In conclusions, this review underlines that many parameters have to be taken into account for the in vivo study of L. donovani complex or L. major infection in mouse models of VL and CL.
From the host genetic point of view, it seems more complex controlling CL than VL. Indeed, several loci are involved in CL control and each locus regulates specific features of the disease (i.e., IgG secretion, parasite load, lesion size). Although the use of an animal model limits the influence of the environment, the choice of the mouse genetic background is crucial. For instance, a sensitive mouse strain is more suitable for comparing the infection outcome of various Leishmania strains and for rapid assessment of the parasite virulence and/or pathogenicity.
Concerning the immune response (resistance/susceptibility phenotype), in VL, mice are clearly classified according to their capacity to sustain parasite persistence in organs. In CL, parasites could persist at the infection site in resistant mice, thus giving a lifelong immunity to reinfection.
For both CL and VL, the parasite genotype clearly affects the infection outcome; however, it is difficult to correlate the results obtained in animal models with clinical observations in human patients.
The parasite dose and the route of inoculation need also to be carefully considered.
Inoculation through the intradermal route must be performed with high dose of parasite inoculum to give effective viscera infection during VL, whereas low doses can be used for CL with parasites dissemination to visceral organs in susceptible mice. In the case of L. infantum infection, different immune responses are observed in function of the inoculation route and this can strongly influence the outcome. Thus, to study the chronic phase of the infection, an inoculation route that promotes the parasite long-term establishment (intravenous route) must be preferred. Conversely, when studying the acute phase, the inoculated dose (high doses leading to more intense immune response) is the crucial point. For L. major , the key parameters seem to be the dose and its association with sandfly saliva. Low doses are recommended for immunization studies, but not for strain virulence assessment. Indeed, at low doses, lesion development is controlled in susceptible mice, except when saliva is simultaneously injected. Effective lesion development requires inoculation of high doses, leading to the development of ulcers the severity of which is directly correlated with the strain virulence.
Abbreviations
Interferon gamma
Interleukin
Leishmania major resistance
Natural killer
Nitric oxyde
Quantitative Trait Locus
Susceptibility to cutaneous leishmaniasis
Transforming growth factor
Tumor necrosis factor
Regulatory T cells
Visceral Leishmaniasis
World Health Organization
Alvar J, Vélez ID, Bern C, Herrero M, Desjeux P, Cano J, Jannin J, de Boer M: Leishmaniasis worldwide and global estimates of its incidence. PLoS ONE 2012;2012;7:1-12.
Bañuls AL, Hide M, Prugnolle F. Leishmania and the Leishmaniases: a parasite genetic update and advances in taxonomy, epidemiology and pathogenicity in humans. Adv Parasitol. 2007;64:1–109.
Article PubMed Google Scholar
Bañuls a L, Bastien P, Pomares C, Arevalo J, Fisa R, Hide M. Clinical pleiomorphism in human leishmaniases, with special mention of asymptomatic infection. Clin Microbiol Infect. 2011;17:1451–61.
Desjeux P. Leishmaniasis: Current situation and new perspectives. Comp Immunol Microbiol Infect Dis. 2004;27:305–18.
Article CAS PubMed Google Scholar
Le Fichoux Y, Quaranta JF, Aufeuvre JP, Lelievre A, Marty P, Suffia I, Rousseau D, Kubar J. Occurrence of Leishmania infantum parasitemia in asymptomatic blood donors living in an area of endemicity in southern France. J Clin Microbiol. 1999;37:1953–7.
Hide M, Marion E, Pomares C, Fisa R, Marty P, Bañuls a L. Parasitic genotypes appear to differ in leishmaniasis patients compared with asymptomatic related carriers. Int J Parasitol. 2013;43:389–97.
Millán J, Ferroglio E, Solano-Gallego L. Role of wildlife in the epidemiology of Leishmania infantum infection in Europe. Parasitol Res. 2014;113(6):2005–14.
Ghawar W, Toumi A, Snoussi M-A, Chlif S, Zâatour A, Boukthir A, Hamida NBH, Chemkhi J, Diouani MF, Ben-Salah A. Leishmania major infection among Psammomys obesus and Meriones shawi : reservoirs of zoonotic cutaneous leishmaniasis in Sidi Bouzid (central Tunisia). Vector Borne Zoonotic Dis. 2011;11:1561–8.
Singh N, Mishra BB, Bajpai S, Singh RK, Tiwari VK. Natural product based leads to fight against leishmaniasis. Bioorg Med Chem. 2014;22(1):18–45.
Larréché S, Launay G, Weibel Galluzzo C, Bousquet A, Eperon G, Pilo J-E, Ravel C, Chappuis F, Dupin M, Mérens A. Cluster of zoonotic cutaneous leishmaniasis ( Leishmania major ) in European travelers returning from Turkmenistan. J Travel Med. 2013;20:400–2.
Mougneau E, Bihl F, Glaichenhaus N. Cell biology and immunology of Leishmania . Immunol Rev. 2011;240:286–96.
Blackwell JM, Fakiola M, Ibrahim ME, Jamieson SE, Jeronimo SB, Miller EN, Mishra A, Mohamed HS, Peacock CS, Raju M, Sundar S, Wilson ME. Genetics and visceral leishmaniasis: Of mice and man. Parasite Immunol. 2009;31(5):254–66.
Vidal S, Tremblay ML, Govoni G, Gauthier S, Sebastiani G, Malo D, Skamene E, Olivier M, Jothy S, Gros P. The Ity/Lsh/Bcg locus: natural resistance to infection with intracellular parasites is abrogated by disruption of the Nramp1 gene. J Exp Med. 1995;182:655–66.
Bradley DJ, Taylor B a, Blackwell J, Evans EP, Freeman J. Regulation of Leishmania populations within the host. III. Mapping of the locus controlling susceptibility to visceral leishmaniasis in the mouse. Clin Exp Immunol. 1979;37:7–14.
CAS PubMed PubMed Central Google Scholar
Kaye PM, Svensson M, Ato M, Maroof A, Polley R, Stager S, Zubairi S, Engwerda CR. The immunopathology of experimental visceral leishmaniasis. Immunol Rev. 2004;201:239–53.
Blackwell JM. Leishmania donovani infection in heterozygous and recombinant H-2 haplotype mice. Immunogenetics. 1983;18:101–9.
Leclercq V, Lebastard M, Belkaid Y, Louis J, Milon G. The outcome of the parasitic process initiated by Leishmania infantum in laboratory mice: a tissue-dependent pattern controlled by the Lsh and MHC loci. J Immunol. 1996;157:4537–45.
CAS PubMed Google Scholar
Engwerda CR, Kaye PM. Organ-specific immune responses associated with infectious disease. Immunol Today. 2000;21:73–8.
Melby PC, Tabares a, Restrepo BI, Cardona a E, McGuff HS, Teale JM. Leishmania donovani : evolution and architecture of the splenic cellular immune response related to control of infection. Exp Parasitol. 2001;99:17–25.
Smelt SC, Engwerda CR, McCrossen M, Kaye PM. Destruction of follicular dendritic cells during chronic visceral leishmaniasis. J Immunol. 1997;158:3813–21.
Murray HW, Stern JJ, Welte K, Carriero SM, Nathan a NDCF, Rubin Y. Treatment With Interleukin 2 and Interferon- $. J Immunol. 1987;138:2290–7.
Kaye PM, Curry a J, Blackwell JM. Differential production of Th1- and Th2-derived cytokines does not determine the genetically controlled or vaccine-induced rate of cure in murine visceral leishmaniasis. J Immunol. 1991;146:2763–70.
Nascimento MSL, Carregaro V, Lima-Júnior DS, Costa DL, Ryffel B, Duthie MS, de Jesus A, de Almeida RP, da Silva JS. Interleukin 17A acts synergistically with interferon γ to promote protection against Leishmania infantum infection. J Infect Dis. 2015;211:1015–26.
Rodrigues OR, Marques C, Soares-Clemente M, Ferronha MH, Santos-Gomes GM. Identification of regulatory T cells during experimental Leishmania infantum infection. Immunobiology. 2009;214:101–11.
Gomes-Pereira S, Rodrigues OR, Rolão N, Almeida PD, Santos-Gomes GM. Hepatic cellular immune responses in mice with “cure” and “non-cure” phenotype to Leishmania infantum infection: Importance of CD8+ T cells and TGF-β production. FEMS Immunol Med Microbiol. 2004;41:59–68.
Squires KE, Kirsch M, Silverstein SC, Acosta A, McElrath MJ, Murray HW. Defect in the tissue cellular immune response: Experimental visceral leishmaniasis in euthymic C57BL/6 ep/ep mice. Infect Immun. 1990;58:3893–8.
Sulahian A, Garin YJF, Pratlong F, Dedet JP, Derouin F. Experimental pathogenicity of viscerotropic and dermotropic isolates of Leishmania infantum from immunocompromised and immunocompetent patients in a murine model. FEMS Immunol Med Microbiol. 1997;17:131–8.
Honoré S, Garin YJF, Sulahian A, Gangneux JP, Derouin F. Influence of the host and parasite strain in a mouse model of visceral Leishmania infantum infection. FEMS Immunol Med Microbiol. 1998;21:231–9.
Gangneux JP, Sulahian A, Honore S, Meneceur P, Derouin F, Garin YJF. Evidence for determining parasitic factors in addition to host genetics and immune status in the outcome of murine Leishmania infantum visceral leishmaniasis. Parasite Immunol. 2000;22:515–9.
Baptista-Fernandes T, Marques C, Roos Rodrigues O, Santos-Gomes GM. Intra-specific variability of virulence in Leishmania infantum zymodeme MON-1 strains. Comp Immunol Microbiol Infect Dis. 2007;30:41–53.
Garin YJF, Sulahian a., Pratlong F, Meneceur P, Gangneux JP, Prina E, Dedet JP, Derouin F. Virulence of Leishmania infantum is expressed as a clonal and dominant phenotype in experimental infections. Infect Immun. 2001;69:7365–73.
Rosypal AC, Zajac AM, Troy GC, Lindsay DS. Infections in immunocompetent and immune-deficient mice with promastigotes of a North American isolate of Leishmania infantum . Vet Parasitol. 2005;130:19–27.
Oliveira DM, Costa MAF, Chavez-Fumagalli M a., Valadares DG, Duarte MC, Costa LE, Martins VT, Gomes RF, Melo MN, Soto M, Tavares CAP, Coelho EAF. Evaluation of parasitological and immunological parameters of Leishmania chagas i infection in BALB/c mice using different doses and routes of inoculation of parasites. Parasitol Res. 2012;110:1277–85.
Kaur S, Kaur T, Garg N, Mukherjee S, Raina P, Athokpam V. Effect of dose and route of inoculation on the generation of CD4 + Th1/Th2 type of immune response in murine visceral leishmaniasis. Parasitol Res. 2008;103:1413–9.
Mukherjee P, Ghosh AK, Ghose AC. Infection pattern and immune response in the spleen and liver of BALB/c mice intracardially infected with Leishmania donovani amastigotes. Immunol Lett. 2003;86:131–8.
Carrión J, Nieto a., Iborra S, Iniesta V, Soto M, Folgueira C, Abanades DR, Requena JM, Alonso C. Immunohistological features of visceral leishmaniasis in BALB/c mice. Parasite Immunol. 2006;28:173–83.
Rolão N, Melo C, Campino L. Influence of the inoculation route in BALB/c mice infected by Leishmania infantum . Acta Trop. 2004;90:123–6.
Ahmed S, Colmenares M, Soong L, Goldsmith-Pestana K, Munstermann L, Molina R, McMahon-Pratt D. Intradermal infection model for pathogenesis and vaccine studies of murine visceral leishmaniasis. Infect Immun. 2003;71:401–10.
Oliveira F, de Carvalho AM, de Oliveira CI. Sand-fly saliva- Leishmania -man: The trigger trio. Front Immunol. 2013;4(NOV):1–8.
Google Scholar
Paranhos M, dos Santos WC, Sherlock I, Oliveira GG, de Carvalho LC. Development of eosinophilia in dogs intradermically inoculated with sand fly saliva and Leishmania ( Leishmania ) chagasi stationary-phase promastigotes. Mem Inst Oswaldo Cruz. 1993;88:249–51.
Dedet JP, Pratlong F, Lanotte G, Ravel C. Cutaneous leishmaniasis. The parasite. Clin Dermatol. 1999;17:261–8.
DeTolla LJ, Scott PA, Farrell JP. Single gene control of resistance to cutaneous leishmaniasis in mice. Immunogenetics. 1981;14:29–39.
Fortier AH, Meltzer MS, Nacy CA. Susceptibility of inbred mice to Leishmania tropica infection: genetic control of the development of cutaneous lesions in P/J mice. J Immunol. 1984;133:454–9.
Mock B a, Fortier a H, Potter M, Nacy C a. Genetic control of systemic Leishmania major infections: Dissociation of intrahepatic amastigote replication from control by the Lsh gene. Infect Immun. 1985;50:588–91.
Davies EV, Singleton AMT, Blackwell JM. Differences in Lsh gene control over systemic Leishmania major and Leishmania donovani or Leishmania mexicana infections are caused by differential targeting to infiltrating and resident liver macrophage populations. Infect Immun. 1988;56:1128–34.
Howard JG, Hale C, Chan-Liew WL. Immunological regulation of experimental cutaneous leishmaniasis. 1. Immunogenetic aspects of susceptibility to Leishmania tropica in mice. Parasite Immunol. 1980;2:303–14.
Mitchell GF, Curtis JM, Handman E. Resistance to cutaneous leishmaniasis in genetically susceptible BALB/c mice. Aust J Exp Biol Med Sci. 1981;59(Pt 5):555–65.
Roberts M, Alexander J, Blackwell JM. Influence of Lsh, H-2, and an H-11-linked gene on visceralization and metastasis associated with Leishmania mexicana infection in mice. Infect Immun. 1989;57:875–81.
Blackwell JM. Genetic susceptibility to leishmanial infections: studies in mice and man. Parasitology. 1996;112(Suppl):S67–74.
PubMed Google Scholar
Foote SJ, Handman E. Genetics of murine leishmaniasis. Brief Funct Genomic Proteomic. 2005;4:270–6.
Sakthianandeswaren A, Foote SJ, Handman E. The role of host genetics in leishmaniasis. Trends Parasitol. 2009;25:383–91.
Kurey I, Kobets T, Havelková H, Slapničková M, Quan L, Trtková K, Grekov I, Svobodová M, Stassen AP, Hutson A, Demant P, Lipoldová M. Distinct genetic control of parasite elimination, dissemination, and disease after Leishmania major infection. Immunogenetics. 2009;61:619–33.
Sacks D, Noben-Trauth N. The immunology of susceptibility and resistance to Leishmania major in mice. Nat Rev Immunol. 2002;2:845–58.
Belkaid Y, Piccirillo C a, Mendez S. CD4 1 CD25 1 regulatory T cells control Leishmania major persistence and immunity. Nature. 2002;420(September):633–7.
Belkaid Y. The role of CD4(+)CD25(+) regulatory T cells in Leishmania infection. Expert Opin Biol Ther. 2003;3:875–85.
Lopez Kostka S, Dinges S, Griewank K, Iwakura Y, Udey MC, von Stebut E. IL-17 promotes progression of cutaneous leishmaniasis in susceptible mice. J Immunol. 2009;182:3039–46.
Lima HC, Titus RG. Effects of sand fly vector saliva on development of cutaneous lesions and the immune response to Leishmania braziliensis in BALB/c mice. Infect Immun. 1996;64:5442–5.
Belkaid Y, Mendez S, Lira R, Kadambi N, Milon G, Sacks D. A natural model of Leishmania major infection reveals a prolonged “silent” phase of parasite amplification in the skin before the onset of lesion formation and immunity. J Immunol. 2000;165:969–77.
Mcelrath BMJ, Kaplan G, Nusrat A, Cohn ZA: The Defect in T Cell Influx in BALB/c Mice. J Exp Med. 1987;165:546-59.
Cangussú SD, de Souza CC, Campos CF, Vieira LQ, Afonso LCC, Arantes RME. Histopathology of Leishmania major infection: Revisiting L. major histopathology in the ear dermis infection model. Mem Inst Oswaldo Cruz. 2009;104:918–22.
Müller K, van Zandbergen G, Hansen B, Laufs H, Jahnke N, Solbach W, Laskay T. Chemokines, natural killer cells and granulocytes in the early course of Leishmania major infection in mice. Med Microbiol Immunol. 2001;190:73–6.
Li J, Nolan TJ, Farrell JP. Leishmania major : a clone with low virulence for BALB/c mice elicits a Th1 type response and protects against infection with a highly virulent clone. Exp Parasitol. 1997;87:47–57.
Kébaïer C, Louzir H, Chenik M, Ben Salah A, Dellagi K. Heterogeneity of wild Leishmania major isolates in experimental murine pathogenicity and specific immune response. Infect Immun. 2001;69:4906–15.
Article PubMed PubMed Central Google Scholar
Markikou-Ouni W, Ben Achour-Chenik Y, Meddeb-Garnaoui a. Effects of Leishmania majo r clones showing different levels of virulence on infectivity, differentiation and maturation of human dendritic cells. Clin Exp Immunol. 2012;169:273–80.
Article CAS PubMed PubMed Central Google Scholar
Alimohammadian MH, Darabi H, Ajdary S, Khaze V, Torkabadi E. Genotypically distinct strains of Leishmania major display diverse clinical and immunological patterns in BALB/c mice. Infect Genet Evol. 2010;10:969–75.
Asadpour A, Riazi-Rad F, Khaze V, Ajdary S, Alimohammadian MH. Distinct strains of Leishmania major induce different cytokine mRNA expression in draining lymph node of BALB/c mice. Parasite Immunol. 2013;35:42–50.
Menon JN, Bretscher P a. Parasite dose determines the Th1/Th2 nature of the response to Leishmania major independently of infection route and strain of host or parasite. Eur J Immunol. 1998;28:4020–8.
Bretscher PA, Wei G, Menon JN, Bielefeldt-Ohmann H. Establishment of stable, cell-mediated immunity that makes “susceptible” mice resistant to Leishmania major . Science. 1992;257:539–42.
Uzonna JE, Wei G, Yurkowski D, Bretscher P. Immune elimination of Leishmania major in mice: implications for immune memory, vaccination, and reactivation disease. J Immunol. 2001;167:6967–74.
Doherty TM, Coffman RL. Leishmania major : effect of infectious dose on T cell subset development in BALB/c mice. Exp Parasitol. 1996;84:124–35.
Uzonna JE, Joyce KL, Scott P. Low dose Leishmania major promotes a transient T helper cell type 2 response that is down-regulated by interferon gamma-producing CD8+ T cells. J Exp Med. 2004;199:1559–66.
Lira R, Doherty M, Modi G, Sacks D. Evolution of lesion formation, parasitic load, immune response, and reservoir potential in C57BL/6 mice following high- and low-dose challenge with Leishmania major . Infect Immun. 2000;68:5176–82.
Nabors GS, Farrell JP. Site-specific immunity to Leishmania major in SWR mice: The site of infection influences susceptibility and expression of the antileishmanial immune response. Infect Immun. 1994;62:3655–62.
Baldwin TM, Elso C, Curtis J, Buckingham L, Handman E: The Site of Leishmania major Infection Determines Disease Severity and Immune Responses. Infect Immun. 2003;71:1–6.
Titus RG, Ribeiro JM. Salivary gland lysates from the sand fly Lutzomyia longipalpis enhance Leishmania infectivity. Science. 1988;239:1306–8.
Theodos CM, Ribeiro JMC, Titus RG. Analysis of enhancing effect of sand fly saliva on Leishmania infection in mice. Infect Immun. 1991;59:1592–8.
Norsworthy NB, Sun J, Elnaiem D, Lanzaro G, Soong L. Sand fly saliva enhances Leishmania amazonensis infection by modulating interleukin-10 production. Infect Immun. 2004;72:1240–7.
Carregaro V, Costa DL, Brodskyn C, Barral AM, Barral-Netto M, Cunha FQ, Silva JS. Dual effect of Lutzomyia longipalpis saliva on Leishmania braziliensis infection is mediated by distinct saliva-induced cellular recruitment into BALB/c mice ear. BMC Microbiol. 2013;13:102.
Morris RV, Shoemaker CB, David JR, Lanzaro GC, Titus RG. Sandfly maxadilan exacerbates infection with Leishmania major and vaccinating against it protects against L. major infection. J Immunol. 2001;167:5226–30.
Mbow ML, Bleyenberg JA, Hall LR, Titus RG. Phlebotomus papatasi sand fly salivary gland lysate down-regulates a Th1, but up-regulates a Th2, response in mice infected with Leishmania major . J Immunol. 1998;161:5571–7.
Belkaid Y, Kamhawi S, Modi G, Valenzuela J, Noben-Trauth N, Rowton E, Ribeiro J, Sacks DL. Development of a natural model of cutaneous leishmaniasis: powerful effects of vector saliva and saliva preexposure on the long-term outcome of Leishmania major infection in the mouse ear dermis. J Exp Med. 1998;188:1941–53.
Lang T, Courret N, Colle JH, Milon G, Antoine JC. The levels and patterns of cytokines produced by CD4 T lymphocytes of BALB/c mice infected with Leishmania major by inoculation into the ear dermis depend on the infectiousness and size of the inoculum. Infect Immun. 2003;71:2674–83.
van Zandbergen G, Bollinger A, Wenzel A, Kamhawi S, Voll R, Klinger M, Müller A, Hölscher C, Herrmann M, Sacks D, Solbach W, Laskay T. Leishmania disease development depends on the presence of apoptotic promastigotes in the virulent inoculum. Proc Natl Acad Sci U S A. 2006;103:13837–42.
Download references
Acknowledgements
This work was supported by the University of Montpellier, UMR IRD 224-CNRS 5290. The authors acknowledge the assistance of Dr E. Andermarcher for English manuscript corrections.
Author information
Authors and affiliations.
Maladies Infectieuses et Vecteurs: Ecologie, Génétique, Evolution et Contrôle (MIVEGEC), UMR IRD 224-CNRS 5290, Institut de Recherche pour le Développement (IRD), Centre National de la Recherche Scientifique (CNRS), Université de Montpellier, Montpellier, France
Corinne Loeuillet, Anne-Laure Bañuls & Mallorie Hide
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Mallorie Hide .
Additional information
Competing interests.
The authors declare that they have no competing interests.
Authors’ contributions
CL reviewed the literature, wrote the paper and made the tables. MH and ALB afforded additional references and critically revised the manuscript. All authors read and approved the final manuscript.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License ( http://creativecommons.org/licenses/by/4.0/ ), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver ( http://creativecommons.org/publicdomain/zero/1.0/ ) applies to the data made available in this article, unless otherwise stated.
Reprints and permissions
About this article
Cite this article.
Loeuillet, C., Bañuls, AL. & Hide, M. Study of Leishmania pathogenesis in mice: experimental considerations. Parasites Vectors 9 , 144 (2016). https://doi.org/10.1186/s13071-016-1413-9
Download citation
Received : 01 October 2015
Accepted : 26 February 2016
Published : 11 March 2016
DOI : https://doi.org/10.1186/s13071-016-1413-9
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Leishmaniases
Parasites & Vectors
ISSN: 1756-3305
- Submission enquiries: [email protected]
Information
- Author Services
Initiatives
You are accessing a machine-readable page. In order to be human-readable, please install an RSS reader.
All articles published by MDPI are made immediately available worldwide under an open access license. No special permission is required to reuse all or part of the article published by MDPI, including figures and tables. For articles published under an open access Creative Common CC BY license, any part of the article may be reused without permission provided that the original article is clearly cited. For more information, please refer to https://www.mdpi.com/openaccess .
Feature papers represent the most advanced research with significant potential for high impact in the field. A Feature Paper should be a substantial original Article that involves several techniques or approaches, provides an outlook for future research directions and describes possible research applications.
Feature papers are submitted upon individual invitation or recommendation by the scientific editors and must receive positive feedback from the reviewers.
Editor’s Choice articles are based on recommendations by the scientific editors of MDPI journals from around the world. Editors select a small number of articles recently published in the journal that they believe will be particularly interesting to readers, or important in the respective research area. The aim is to provide a snapshot of some of the most exciting work published in the various research areas of the journal.
Original Submission Date Received: .
- Active Journals
- Find a Journal
- Proceedings Series
- For Authors
- For Reviewers
- For Editors
- For Librarians
- For Publishers
- For Societies
- For Conference Organizers
- Open Access Policy
- Institutional Open Access Program
- Special Issues Guidelines
- Editorial Process
- Research and Publication Ethics
- Article Processing Charges
- Testimonials
- Preprints.org
- SciProfiles
- Encyclopedia

Article Menu

- Subscribe SciFeed
- Google Scholar
- on Google Scholar
- Table of Contents
Find support for a specific problem in the support section of our website.
Please let us know what you think of our products and services.
Visit our dedicated information section to learn more about MDPI.
JSmol Viewer
The state-of-the-art of mycobacterium chimaera infections and the causal link with health settings: a systematic review.

1. Introduction
2. materials and methods, 4. discussion, 4.1. mycobacterium chimaera’s characteristics and ecosystem, 4.2. heater-cooler units, medical devices, water, and air-conditioned implants, 4.3. incubation period and symptoms presentation, 4.4. presence in the lung system, 4.5. modality of transmission, 4.6. detection, 4.7. disinfection, 4.8. causal link assessment, 5. limitations, 6. conclusions, supplementary materials, author contributions, institutional review board statement, informed consent statement, data availability statement, conflicts of interest, abbreviations.
| MAC | mycobacterium avium complex |
| NTM | non-tuberculosis mycobacterium |
| M. chimaera | Mycobacterium chimaera |
| HCU | heater-cooler units |
| OPPP | opportunistic premise plumbing pathogens |
| ECMO | extra-corporal mechanical oxygenation |
| HAI | healthcare-associated infection |
- Vendramin, I.; Peghin, M.; Tascini, C.; Livi, U. Longest Incubation Period of Mycobacterium chimaera Infection after Cardiac Surgery. Eur. J. Cardio-Thoracic Surg. 2021 , 59 , 506–508. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Biswas, M.; Rahaman, S.; Biswas, T.K.; Haque, Z.; Ibrahim, B. Association of Sex, Age, and Comorbidities with Mortality in COVID-19 Patients: A Systematic Review and Meta-Analysis. Intervirology 2020 , 64 , 36–47. [ Google Scholar ] [ CrossRef ]
- Treglia, M.; Pallocci, M.; Passalacqua, P.; Sabatelli, G.; De Luca, L.; Zanovello, C.; Messineo, A.; Quintavalle, G.; Cisterna, A.M.; Marsella, L.T. Medico-Legal Aspects of Hospital-Acquired Infections: 5-Years of Judgements of the Civil Court of Rome. Healthcare 2022 , 10 , 1336. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bolcato, V.; Tronconi, L.P.; Odone, A.; Blandi, L. Healthcare-Acquired Sars-Cov-2 Infection: A Viable Legal Category? Int. J. Risk Saf. Med. 2023 , 34 , 129–134. [ Google Scholar ] [ CrossRef ]
- Tattoli, L.; Dell’erba, A.; Ferorelli, D.; Gasbarro, A.; Solarino, B. Sepsis and Nosocomial Infections: The Role of Medico-Legal Experts in Italy. Antibiotics 2019 , 8 , 199. [ Google Scholar ] [ CrossRef ]
- Barranco, R.; Caristo, I.; Spigno, F.; Ponzano, M.; Trevisan, A.; Signori, A.; Di Biagio, A.; Ventura, F. Management of the Medico-Legal Dispute of Healthcare-Related SARS-CoV-2 Infections: Evaluation Criteria and Case Study in a Large University Hospital in Northwest Italy from 2020 to 2021. Int. J. Environ. Res. Public Health 2022 , 19 , 16764. [ Google Scholar ] [ CrossRef ]
- Goldenberg, S.D.; Volpé, H.; French, G.L. Clinical Negligence, Litigation and Healthcare-Associated Infections. J. Hosp. Infect. 2012 , 81 , 156–162. [ Google Scholar ] [ CrossRef ]
- Rizzo, N. La Causalità Civile ; Jus Civile; Giappichelli Editore: Torino, Italy, 2022. [ Google Scholar ]
- Riccardi, N.; Monticelli, J.; Antonello, R.M.; Luzzati, R.; Gabrielli, M.; Ferrarese, M.; Codecasa, L.; Di Bella, S.; Giacobbe, D.R. Mycobacterium chimaera Infections: An Update. J. Infect. Chemother. 2020 , 26 , 199–205. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Natanti, A.; Palpacelli, M.; Valsecchi, M.; Tagliabracci, A.; Pesaresi, M. Mycobacterium chimaera : A Report of 2 New Cases and Literature Review. Int. J. Leg. Med. 2021 , 135 , 2667–2679. [ Google Scholar ] [ CrossRef ]
- Wetzstein, N.; Kohl, T.A.; Diricks, M.; Mas-Peiro, S.; Holubec, T.; Kessel, J.; Graf, C.; Koch, B.; Herrmann, E.; Vehreschild, M.J.G.T.; et al. Clinical Characteristics and Outcome of Mycobacterium chimaera Infections after Cardiac Surgery: Systematic Review and Meta-Analysis of 180 Heater-Cooler Unit-Associated Cases. Clin. Microbiol. Infect. 2023 , 29 , 1008–1014. [ Google Scholar ] [ CrossRef ]
- Desai, A.N.; Hurtado, R.M. Infections and Outbreaks of Nontuberculous Mycobacteria in Hospital Settings. Curr. Treat. Options Infect. Dis. 2018 , 10 , 169–181. [ Google Scholar ] [ CrossRef ]
- van Ingen, J.; Kohl, T.A.; Kranzer, K.; Hasse, B.; Keller, P.M.; Katarzyna Szafrańska, A.; Hillemann, D.; Chand, M.; Schreiber, P.W.; Sommerstein, R.; et al. Global Outbreak of Severe Mycobacterium chimaera Disease after Cardiac Surgery: A Molecular Epidemiological Study. Lancet Infect. Dis. 2017 , 17 , 1033–1041. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bisognin, F.; Messina, F.; Butera, O.; Nisii, C.; Mazzarelli, A.; Cristino, S.; Pascale, M.R.; Lombardi, G.; Cannas, A.; Dal Monte, P. Investigating the Origin of Mycobacterium chimaera Contamination in Heater-Cooler Units: Integrated Analysis with Fourier Transform Infrared Spectroscopy and Whole-Genome Sequencing. Microbiol. Spectr. 2022 , 10 , e0289322. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Pinzauti, D.; De Giorgi, S.; Fox, V.; Lazzeri, E.; Messina, G.; Santoro, F.; Iannelli, F.; Ricci, S.; Pozzi, G. Complete Genome Sequences of Mycobacterium chimaera Strains 850 and 852, Isolated from Heater-Cooler Unit Water. Microbiol. Resour. Announc. 2022 , 11 , e0102121. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Hasan, N.A.; Warren, R.L.; Elaine Epperson, L.; Malecha, A.; Alexander, D.C.; Turenne, C.Y.; MacMillan, D.; Birol, I.; Pleasance, S.; Coope, R.; et al. Complete Genome Sequence of Mycobacterium chimaera SJ42, a Nonoutbreak Strain from an Immunocompromised Patient with Pulmonary Disease. Genome Announc. 2017 , 5 , e00963-17. [ Google Scholar ] [ CrossRef ]
- Wallace, R.J.; Iakhiaeva, E.; Williams, M.D.; Brown-Elliott, B.A.; Vasireddy, S.; Vasireddy, R.; Lande, L.; Peterson, D.D.; Sawicki, J.; Kwait, R.; et al. Absence of Mycobacterium Intracellulare and Presence of Mycobacterium chimaera in Household Water and Biofilm Samples of Patients in the United States with Mycobacterium avium Complex Respiratory Disease. J. Clin. Microbiol. 2013 , 51 , 1747–1752. [ Google Scholar ] [ CrossRef ]
- Falkinham, J.O.; Hilborn, E.D.; Arduino, M.J.; Pruden, A.; Edwards, M.A. Epidemiology and Ecology of Opportunistic Premise Plumbing Pathogens: Legionella pneumophila , Mycobacterium avium , and Pseudomonas aeruginosa . Environ. Health Perspect. 2015 , 123 , 749–758. [ Google Scholar ] [ CrossRef ]
- European Centre for Disease Prevention and Control. EU Protocol for Testing of M. chimaera Infections Potentially Associated with Heater-Cooler Units Environmental Microbiology Investigations ; Technical Document; European Centre for Disease Prevention and Control: Solna, Sweden, 2015. [ Google Scholar ]
- Ministero della Salute. Raccomandazioni per Il Controllo Dell’infezione da Mycobacterium chimaera in Italia ; Ministero della Salute: Roma, Italy, 2019.
- Bolcato, M.; Rodriguez, D.; Aprile, A. Risk Management in the New Frontier of Professional Liability for Nosocomial Infection: Review of the Literature on Mycobacterium chimaera . Int. J. Environ. Res. Public Health 2020 , 17 , 7328. [ Google Scholar ] [ CrossRef ]
- Achermann, Y.; Rössle, M.; Hoffmann, M.; Deggim, V.; Kuster, S.; Zimmermann, D.R.; Bloemberg, G.; Hombach, M.; Hasse, B. Prosthetic Valve Endocarditis and Bloodstream Infection Due to Mycobacterium chimaera . J. Clin. Microbiol. 2013 , 51 , 1769–1773. [ Google Scholar ] [ CrossRef ]
- Zabost, A.T.; Szturmowicz, M.; Brzezińska, S.A.; Klatt, M.D.; Augustynowicz-Kopeć, E.M. Mycobacterium chimaera as an Underestimated Cause of NTM Lung Diseases in Patients Hospitalized in Pulmonary Wards. Pol. J. Microbiol. 2021 , 70 , 315–320. [ Google Scholar ] [ CrossRef ]
- Truden, S.; Žolnir-Dovč, M.; Sodja, E.; Starčič Erjavec, M. Nationwide Analysis of Mycobacterium chimaera and Mycobacterium intracellulare Isolates: Frequency, Clinical Importance, and Molecular and Phenotypic Resistance Profiles. Infect. Genet. Evol. 2020 , 82 , 104311. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 Statement: An Updated Guideline for Reporting Systematic Reviews. BMJ 2021 , 372 , 71. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bills, N.D.; Hinrichs, S.H.; Aden, T.A.; Wickert, R.S.; Iwen, P.C. Molecular Identification of Mycobacterium chimaera as a Cause of Infection in a Patient with Chronic Obstructive Pulmonary Disease. Diagn. Microbiol. Infect. Dis. 2009 , 63 , 292–295. [ Google Scholar ] [ CrossRef ]
- Cohen-Bacrie, S.; David, M.; Stremler, N.; Dubus, J.-C.; Rolain, J.-M.; Drancourt, M. Mycobacterium chimaera Pulmonary Infection Complicating Cystic Fibrosis: A Case Report. J. Med. Case Rep. 2011 , 5 , 473. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Alhanna, J.; Purucker, M.; Steppert, C.; Grigull-Daborn, A.; Schiffel, G.; Gruber, H.; Borgmann, S. Mycobacterium chimaera Causes Tuberculosis-like Infection in a Male Patient with Anorexia Nervosa. Int. J. Eat. Disord. 2012 , 45 , 450–452. [ Google Scholar ] [ CrossRef ]
- Gunaydin, M.; Yanik, K.; Eroglu, C.; Sanic, A.; Ceyhan, I.; Erturan, Z.; Durmaz, R. Distribution of Nontuberculous Mycobacteria Strains. Ann. Clin. Microbiol. Antimicrob. 2013 , 12 , 33. [ Google Scholar ] [ CrossRef ]
- Boyle, D.P.; Zembower, T.R.; Reddy, S.; Qi, C. Comparison of Clinical Features, Virulence, and Relapse among Mycobacterium avium Complex Species. Am. J. Respir. Crit. Care Med. 2015 , 191 , 1310–1317. [ Google Scholar ] [ CrossRef ]
- Mwikuma, G.; Kwenda, G.; Hang’ombe, B.M.; Simulundu, E.; Kaile, T.; Nzala, S.; Siziya, S.; Suzuki, Y. Molecular Identification of Non-Tuberculous Mycobacteria Isolated from Clinical Specimens in Zambia. Ann. Clin. Microbiol. Antimicrob. 2015 , 14 , 1. [ Google Scholar ] [ CrossRef ]
- Moon, S.M.; Kim, S.Y.; Jhun, B.W.; Lee, H.; Park, H.Y.; Jeon, K.; Huh, H.J.; Ki, C.S.; Lee, N.Y.; Shin, S.J.; et al. Clinical Characteristics and Treatment Outcomes of Pulmonary Disease Caused by Mycobacterium chimaera . Diagn. Microbiol. Infect. Dis. 2016 , 86 , 382–384. [ Google Scholar ] [ CrossRef ]
- Moutsoglou, D.M.; Merritt, F.; Cumbler, E. Disseminated Mycobacterium chimaera Presenting as Vertebral Osteomyelitis. Case Rep. Infect. Dis. 2017 , 2017 , 9893743. [ Google Scholar ] [ CrossRef ]
- Bursle, E.; Playford, E.G.; Coulter, C.; Griffin, P. First Australian Case of Disseminated Mycobacterium chimaera Infection Post-Cardiothoracic Surgery. Infect. Dis. Health 2017 , 22 , 1–5. [ Google Scholar ] [ CrossRef ]
- Kim, S.-Y.; Shin, S.H.; Moon, S.M.; Yang, B.; Kim, H.; Kwon, O.J.; Huh, H.J.; Ki, C.-S.; Lee, N.Y.; Shin, S.J.; et al. Distribution and Clinical Significance of Mycobacterium avium Complex Species Isolated from Respiratory Specimens. Diagn. Microbiol. Infect. Dis. 2017 , 88 , 125–137. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Chand, M.; Lamagni, T.; Kranzer, K.; Hedge, J.; Moore, G.; Parks, S.; Collins, S.; Del Ojo Elias, C.; Ahmed, N.; Brown, T.; et al. Insidious Risk of Severe Mycobacterium chimaera Infection in Cardiac Surgery Patients. Clin. Infect. Dis. 2017 , 64 , 335–342. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Truden, S.; Žolnir-Dovč, M.; Sodja, E.; Starčič Erjavec, M. Retrospective Analysis of Slovenian Mycobacterium avium Complex and Mycobacterium abscessus Complex Isolates and Molecular Resistance Profile. Russ. J. Infect. Immun. 2018 , 8 , 447–451. [ Google Scholar ] [ CrossRef ]
- Larcher, R.; Lounnas, M.; Dumont, Y.; Michon, A.L.; Bonzon, L.; Chiron, R.; Carriere, C.; Klouche, K.; Godreuil, S. Mycobacterium chimaera Pulmonary Disease in Cystic Fibrosis Patients, France, 2010–2017. Emerg. Infect. Dis. 2019 , 25 , 611–613. [ Google Scholar ] [ CrossRef ]
- Shafizadeh, N.; Hale, G.; Bhatnagar, J.; Alshak, N.S.; Nomura, J. Mycobacterium chimaera Hepatitis: A New Disease Entity. Am. J. Surg. Pathol. 2019 , 43 , 244–250. [ Google Scholar ] [ CrossRef ]
- Rosero, C.I.; Shams, W.E. Mycobacterium chimaera Infection Masquerading as a Lung Mass in a Healthcare Worker. IDCases 2019 , 15 , e00526. [ Google Scholar ] [ CrossRef ]
- Watanabe, R.; Seino, H.; Taniuchi, S.; Igusa, R. Mycobacterium chimaera -Induced Tenosynovitis in a Patient with Rheumatoid Arthritis. BMJ Case Rep. 2020 , 13 , e233868. [ Google Scholar ] [ CrossRef ]
- Chen, L.C.; Huang, H.N.; Yu, C.J.; Chien, J.Y.; Hsueh, P.R. Clinical Features and Treatment Outcomes of Mycobacterium chimaera Lung Disease and Antimicrobial Susceptibility of the Mycobacterial Isolates. J. Infect. 2020 , 80 , 437–443. [ Google Scholar ] [ CrossRef ]
- Maalouly, C.; Devresse, A.; Martin, A.; Rodriguez-Villalobos, H.; Kanaan, N.; Belkhir, L. Coinfection of Mycobacterium malmoense and Mycobacterium chimaera in a Kidney Transplant Recipient: A Case Report and Review of the Literature. Transpl. Infect. Dis. 2020 , 22 , e13241. [ Google Scholar ] [ CrossRef ]
- de Melo Carvalho, R.; Nunes, A.L.; Sa, R.; Ramos, I.; Valente, C.; Saraiva da Cunha, J. Mycobacterium chimaera Disseminated Infection. J. Med. Cases 2020 , 11 , 35–36. [ Google Scholar ] [ CrossRef ]
- Sharma, K.; Sharma, M.; Modi, M.; Joshi, H.; Goyal, M.; Sharma, A.; Ray, P.; Rowlinson, M.C. Mycobacterium chimaera and Chronic Meningitis. QJM Int. J. Med. 2020 , 113 , 563–564. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Kim, M.J.; Kim, K.M.; Shin, J.I.; Ha, J.H.; Lee, D.H.; Choi, J.G.; Park, J.S.; Byun, J.H.; Yoo, J.W.; Eum, S.; et al. Identification of Nontuberculous Mycobacteria in Patients with Pulmonary Diseases in Gyeongnam, Korea, Using Multiplex PCR and Multigene Sequence-Based Analysis. Can. J. Infect. Dis. Med. Microbiol. 2021 , 2021 , 8844306. [ Google Scholar ] [ CrossRef ]
- Kavvalou, A.; Stehling, F.; Tschiedel, E.; Kehrmann, J.; Walkenfort, B.; Hasenberg, M.; Olivier, M.; Steindor, M. Biofilm Infection of a Central Venous Port-Catheter Caused by Mycobacterium avium Complex in an Immunocompetent Child with Cystic Fibrosis. BMC Infect. Dis. 2022 , 22 , 321. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Robinson, B.; Chaudhri, M.; Miskoff, J.A. A Case of Cavitary Mycobacterium chimaera . Cureus 2022 , 14 , e26984. [ Google Scholar ] [ CrossRef ]
- Ahmad, M.; Yousaf, A.; Khan, H.M.W.; Munir, A.; Chandran, A. Mycobacterium chimaera Lung Infection and Empyema in a Patient without Cardiopulmonary Bypass. Bayl. Univ. Med. Cent. Proc. 2022 , 35 , 817–819. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- George, M.; Afra, T.P.; Santhosh, P.; Nandakumar, G.; Balagopalan, D.; Sreedharan, S. Ulcerating Nodules on the Face Due to Mycobacterium chimaera in a Patient with Diabetes. Clin. Exp. Dermatol. 2022 , 47 , 587–589. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Lin, Y.F.; Lee, T.F.; Wu, U.I.; Huang, C.F.; Cheng, A.; Lin, K.Y.; Hung, C.C. Disseminated Mycobacterium chimaera Infection in a Patient with Adult-Onset Immunodeficiency Syndrome: Case Report. BMC Infect. Dis. 2022 , 22 , 665. [ Google Scholar ] [ CrossRef ]
- Łyżwa, E.; Siemion-Szcześniak, I.; Sobiecka, M.; Lewandowska, K.; Zimna, K.; Bartosiewicz, M.; Jakubowska, L.; Augustynowicz-Kopeć, E.; Tomkowski, W. An Unfavorable Outcome of M. Chimaera Infection in Patient with Silicosis. Diagnostics 2022 , 12 , 1826. [ Google Scholar ] [ CrossRef ]
- McLaughlin, C.M.; Schade, M.; Cochran, E.; Taylor, K.F. A Case Report of a Novel Atypical Mycobacterial Infection: Mycobacterium chimaera Hand Tenosynovitis. JBJS Case Connect. 2022 , 12 , e22. [ Google Scholar ] [ CrossRef ]
- Gross, J.E.; Teneback, C.C.; Sweet, J.G.; Caceres, S.M.; Poch, K.R.; Hasan, N.A.; Jia, F.; Epperson, L.E.; Lipner, E.M.; Vang, C.K.; et al. Molecular Epidemiologic Investigation of Mycobacterium intracellulare Subspecies Chimaera Lung Infections at an Adult Cystic Fibrosis Program. Ann. Am. Thorac. Soc. 2023 , 20 , 677–686. [ Google Scholar ] [ CrossRef ]
- Azzarà, C.; Lombardi, A.; Gramegna, A.; Ori, M.; Gori, A.; Blasi, F.; Bandera, A. Non-Tuberculous Mycobacteria Lung Disease Due to Mycobacterium chimaera in a 67-Year-Old Man Treated with Immune Checkpoint Inhibitors for Lung Adenocarcinoma: Infection Due to Dysregulated Immunity? BMC Infect. Dis. 2023 , 23 , 573. [ Google Scholar ] [ CrossRef ]
- Pradhan, A.; Martinez, E.; Sintchenko, V.; Post, J.; Overton, K. Case of Mycobacterium chimaera Vertebral Osteomyelitis Diagnosed 7 Years after Cardiac Surgery. Intern. Med. J. 2023 , 53 , 150–151. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Garcia-Prieto, F.; Rodríguez Perojo, A.; Río Ramírez, M.T. Endobronchial Fibroanthracosis Associated with Mycobacterium chimaera Infection: An Exceptional Case. Open Respir. Arch. 2024 , 6 , 100309. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Paul, S.; MacNair, A.; Lostarakos, V.; Capstick, R. Non-Tuberculous Mycobacterial Pulmonary Infection Presenting in a Patient with Unilateral Pulmonary Artery Agenesis. BMJ Case Rep. 2024 , 17 , e259125. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bittner, M.J.; Preheim, L.C. Other Slow-Growing Nontuberculous Mycobacteria. Microbiol. Spectr. 2016 , 4 , 767–776. [ Google Scholar ] [ CrossRef ]
- Tortoli, E.; Rindi, L.; Garcia, M.J.; Chiaradonna, P.; Dei, R.; Garzelli, C.; Kroppenstedt, R.M.; Lari, N.; Mattei, R.; Mariottini, A.; et al. Proposal to Elevate the Genetic Variant MAC-A Included in the Mycobacterium avium Complex, to Species Rank as Mycobacterium chimaera sp. nov. Int. J. Syst. Evol. Microbiol. 2004 , 54 , 1277–1285. [ Google Scholar ] [ CrossRef ]
- Turankar, R.P.; Singh, V.; Gupta, H.; Pathak, V.K.; Ahuja, M.; Singh, I.; Lavania, M.; Dinda, A.K.; Sengupta, U. Association of Non-Tuberculous Mycobacteria with Mycobacterium leprae in Environment of Leprosy Endemic Regions in India. Infect. Genet. Evol. 2019 , 72 , 191–198. [ Google Scholar ] [ CrossRef ]
- Makovcova, J.; Slany, M.; Babak, V.; Slana, I.; Kralik, P. The Water Environment as a Source of Potentially Pathogenic Mycobacteria. J. Water Health 2014 , 12 , 254–263. [ Google Scholar ] [ CrossRef ]
- Falkinham, J.O. Ecology of Nontuberculous Mycobacteria-Where Do Human Infections Come From? Semin. Respir. Crit. Care Med. 2013 , 34 , 95–102. [ Google Scholar ] [ CrossRef ]
- Norton, G.J.; Williams, M.; Falkinham, J.O., III; Honda, J.R. Physical Measures to Reduce Exposure to Tap Water-Associated Nontuberculous Mycobacteria. Front. Public Health 2020 , 8 , 190. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Durnez, L.; Eddyani, M.; Mgode, G.F.; Katakweba, A.; Katholi, C.R.; Machang’u, R.R.; Kazwala, R.R.; Portaels, F.; Leirs, H. First Detection of Mycobacteria in African Rodents and Insectivores, Using Stratified Pool Screening. Appl. Environ. Microbiol. 2008 , 74 , 768. [ Google Scholar ] [ CrossRef ]
- Sax, H.; Bloemberg, G.; Hasse, B.; Sommerstein, R.; Kohler, P.; Achermann, Y.; Rössle, M.; Falk, V.; Kuster, S.P.; Böttger, E.C.; et al. Prolonged Outbreak of Mycobacterium chimaera Infection after Open-Chest Heart Surgery. Clin. Infect. Dis. 2015 , 61 , 67–75. [ Google Scholar ] [ CrossRef ]
- Falkinham, J.O.; Williams, M.D. Desiccation-Tolerance of Mycobacterium avium , Mycobacterium intracellulare , Mycobacterium chimaera , Mycobacterium abscessus and Mycobacterium chelonae . Pathogens 2022 , 11 , 463. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Gebert, M.J.; Delgado-Baquerizo, M.; Oliverio, A.M.; Webster, T.M.; Nichols, L.M.; Honda, J.R.; Chan, E.D.; Adjemian, J.; Dunn, R.R.; Fierer, N. Ecological Analyses of Mycobacteria in Showerhead Biofilms and Their Relevance to Human Health. mBio 2018 , 9 , e01614–e01618. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Cao, Y.; Yuan, S.; Pang, L.; Xie, J.; Gao, Y.; Zhang, J.; Zhao, Z.; Yao, S. Study on Microbial Diversity of Washing Machines. Biodegradation 2024 , 1–13. [ Google Scholar ] [ CrossRef ]
- Liu, H.; Jiao, P.; Guan, L.; Wang, C.; Zhang, X.X.; Ma, L. Functional Traits and Health Implications of the Global Household Drinking-Water Microbiome Retrieved Using an Integrative Genome-Centric Approach. Water Res. 2024 , 250 , 121094. [ Google Scholar ] [ CrossRef ]
- Choi, J.Y.; Sim, B.R.; Park, Y.; Yong, S.H.; Shin, S.J.; Kang, Y.A. Identification of Nontuberculous Mycobacteria Isolated from Household Showerheads of Patients with Nontuberculous Mycobacteria. Sci. Rep. 2022 , 12 , 8648. [ Google Scholar ] [ CrossRef ]
- Shen, Y.; Haig, S.J.; Prussin, A.J.; Lipuma, J.J.; Marr, L.C.; Raskin, L. Shower Water Contributes Viable Nontuberculous Mycobacteria to Indoor Air. PNAS Nexus 2022 , 1 , pgac145. [ Google Scholar ] [ CrossRef ]
- Struelens, M.J.; Plachouras, D. Mycobacterium chimaera Infections Associated with Heater-Cooler Units (HCU): Closing Another Loophole in Patient Safety. Eurosurveillance 2016 , 21 , 30397. [ Google Scholar ] [ CrossRef ]
- Trudzinski, F.C.; Schlotthauer, U.; Kamp, A.; Hennemann, K.; Muellenbach, R.M.; Reischl, U.; Gärtner, B.; Wilkens, H.; Bals, R.; Herrmann, M.; et al. Clinical Implications of Mycobacterium chimaera Detection in Thermoregulatory Devices Used for Extracorporeal Membrane Oxygenation (ECMO), Germany, 2015 to 2016. Eurosurveillance 2016 , 21 , 30398. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Schnetzinger, M.; Heger, F.; Indra, A.; Kimberger, O. Bacterial Contamination of Water Used as Thermal Transfer Fluid in Fluid-Warming Devices. J. Hosp. Infect. 2023 , 141 , 49–54. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Barker, T.A.; Dandekar, U.; Fraser, N.; Dawkin, L.; Sweeney, P.; Heron, F.; Simmons, J.; Parmar, J. Minimising the Risk of Mycobacterium chimaera Infection during Cardiopulmonary Bypass by the Removal of Heater-Cooler Units from the Operating Room. Perfusion 2018 , 33 , 264–269. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Barnes, S.; Twomey, C.; Carrico, R.; Murphy, C.; Warye, K. OR Air Quality: Is It Time to Consider Adjunctive Air Cleaning Technology? AORN J. 2018 , 108 , 503–515. [ Google Scholar ] [ CrossRef ]
- Walker, J.T.; Lamagni, T.; Chand, M. Evidence That Mycobacterium chimaera Aerosols Penetrate Laminar Airflow and Result in Infections at the Surgical Field. Lancet Infect. Dis. 2017 , 17 , 1019. [ Google Scholar ] [ CrossRef ]
- Schlotthauer, U.; Hennemann, K.; Gärtner, B.C.; Schäfers, H.-J.; Becker, S.L. Microbiological Surveillance of Heater-Cooler Units Used in Cardiothoracic Surgery for Detection of Mycobacterium chimaera . Thorac. Cardiovasc. Surg. 2022 , 72 , 59–62. [ Google Scholar ] [ CrossRef ]
- Gross, J.E.; Caceres, S.; Poch, K.; Epperson, L.E.; Hasan, N.A.; Jia, F.; de Moura, V.C.N.; Strand, M.; Lipner, E.M.; Honda, J.R.; et al. Prospective Healthcare-Associated Links in Transmission of Nontuberculous Mycobacteria among People with Cystic Fibrosis (PHALT NTM) Study: Rationale and Study Design. PLoS ONE 2023 , 18 , e0291910. [ Google Scholar ] [ CrossRef ]
- Nakamura, S.; Azuma, M.; Sato, M.; Fujiwara, N.; Nishino, S.; Wada, T.; Yoshida, S. Pseudo-Outbreak of Mycobacterium chimaera through Aerators of Hand-Washing Machines at a Hematopoietic Stem Cell Transplantation Center. Infect. Control Hosp. Epidemiol. 2019 , 40 , 1433–1435. [ Google Scholar ] [ CrossRef ]
- Kanamori, H.; Weber, D.J.; Rutala, W.A. Healthcare-Associated Mycobacterium chimaera Transmission and Infection Prevention Challenges: Role of Heater-Cooler Units as a Water Source in Cardiac Surgery. Clin. Infect. Dis. 2017 , 64 , 343–346. [ Google Scholar ] [ CrossRef ]
- Rao, M.; Silveira, F.P. Non-Tuberculous Mycobacterial Infections in Thoracic Transplant Candidates and Recipients. Curr. Infect. Dis. Rep. 2018 , 20 , 14. [ Google Scholar ] [ CrossRef ]
- Walker, J.; Moore, G.; Collins, S.; Parks, S.; Garvey, M.I.; Lamagni, T.; Smith, G.; Dawkin, L.; Goldenberg, S.; Chand, M. Microbiological Problems and Biofilms Associated with Mycobacterium chimaera in Heater–Cooler Units Used for Cardiopulmonary Bypass. J. Hosp. Infect. 2017 , 96 , 209–220. [ Google Scholar ] [ CrossRef ]
- Born, F.; Wieser, A.; Oberbach, A.; Oberbach, A.; Ellgass, R.; Peterss, S.; Kur, F.; Grabein, B.; Hagl, C. Five Years without Mycobacterium chimaera . Thorac. Cardiovasc. Surg. 2020 , 68 , S1–S72. [ Google Scholar ] [ CrossRef ]
- Scriven, J.E.; Scobie, A.; Verlander, N.Q.; Houston, A.; Collyns, T.; Cajic, V.; Kon, O.M.; Mitchell, T.; Rahama, O.; Robinson, A.; et al. Mycobacterium chimaera Infection Following Cardiac Surgery in the United Kingdom: Clinical Features and Outcome of the First 30 Cases. Clin. Microbiol. Infect. 2018 , 24 , 1164–1170. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Olatidoye, O.A.; Samat, S.H.; Yin, K.; Bates, M.J. Pulmonary Valve Infective Endocarditis Caused by Mycobacterium abscessus . J. Cardiothorac. Surg. 2023 , 18 , 221. [ Google Scholar ] [ CrossRef ]
- Ganatra, S.; Sharma, A.; D’Agostino, R.; Gage, T.; Kinnunen, P. Mycobacterium chimaera Mimicking Sarcoidosis. Methodist Debakey Cardiovasc. J. 2018 , 14 , 301–302. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Buchanan, R.; Agarwal, A.; Mathai, E.; Cherian, B. Mycobacterium chimaera : A Novel Pathogen with Potential Risk to Cardiac Surgical Patients. Natl. Med. J. India 2020 , 33 , 284–287. [ Google Scholar ] [ CrossRef ]
- McHugh, J.; Saleh, O.A. Updates in Culture-Negative Endocarditis. Pathogens 2023 , 12 , 1027. [ Google Scholar ] [ CrossRef ]
- Delgado, V.; Ajmone Marsan, N.; De Waha, S.; Bonaros, N.; Brida, M.; Burri, H.; Caselli, S.; Doenst, T.; Ederhy, S.; Erba, P.A.; et al. 2023 ESC Guidelines for the Management of Endocarditis. Eur. Heart J. 2023 , 44 , 3948–4042. [ Google Scholar ] [ CrossRef ]
- Kohler, P.; Kuster, S.P.; Bloemberg, G.; Schulthess, B.; Frank, M.; Tanner, F.C.; Rössle, M.; Böni, C.; Falk, V.; Wilhelm, M.J.; et al. Healthcare-Associated Prosthetic Heart Valve, Aortic Vascular Graft, and Disseminated Mycobacterium chimaera Infections Subsequent to Open Heart Surgery. Eur. Heart J. 2015 , 36 , 2745–2753. [ Google Scholar ] [ CrossRef ]
- Wyrostkiewicz, D.; Opoka, L.; Filipczak, D.; Jankowska, E.; Skorupa, W.; Augustynowicz-Kopeć, E.; Szturmowicz, M. Nontuberculous Mycobacterial Lung Disease in the Patients with Cystic Fibrosis—A Challenging Diagnostic Problem. Diagnostics 2022 , 12 , 1514. [ Google Scholar ] [ CrossRef ]
- Virdi, R.; Lowe, M.E.; Norton, G.J.; Dawrs, S.N.; Hasan, N.A.; Epperson, L.E.; Glickman, C.M.; Chan, E.D.; Strong, M.; Crooks, J.L.; et al. Lower Recovery of Nontuberculous Mycobacteria from Outdoor Hawai’i Environmental Water Biofilms Compared to Indoor Samples. Microorganisms 2021 , 9 , 224. [ Google Scholar ] [ CrossRef ]
- Schweickert, B.; Goldenberg, O.; Richter, E.; Göbel, U.B.; Petrich, A.; Buchholz, P.; Moter, A. Occurrence and Clinical Relevance of Mycobacterium chimaera sp. nov., Germany. Emerg. Infect. Dis. 2008 , 14 , 1443–1446. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Boyle, D.P.; Zembower, T.R.; Qi, C. Evaluation of Vitek MS for Rapid Classification of Clinical Isolates Belonging to Mycobacterium avium Complex. Diagn. Microbiol. Infect. Dis. 2015 , 81 , 41–43. [ Google Scholar ] [ CrossRef ]
- Sommerstein, R.; Rüegg, C.; Kohler, P.; Bloemberg, G.; Kuster, S.P.; Sax, H. Transmission of Mycobacterium chimaera from Heater-Cooler Units during Cardiac Surgery despite an Ultraclean Air Ventilation System. Emerg. Infect. Dis. 2016 , 22 , 1008–1013. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Balsam, L.B.; Louie, E.; Hill, F.; Levine, J.; Phillips, M.S. Mycobacterium chimaera Left Ventricular Assist Device Infections. J. Card. Surg. 2017 , 32 , 402–404. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Sanchez-Nadales, A.; Diaz-Sierra, A.; Mocadie, M.; Asher, C.; Gordon, S.; Xu, B. Advanced Cardiovascular Imaging for the Diagnosis of Mycobacterium chimaera Prosthetic Valve Infective Endocarditis After Open-Heart Surgery: A Contemporary Systematic Review. Curr. Probl. Cardiol. 2022 , 47 , 101392. [ Google Scholar ] [ CrossRef ]
- Cannas, A.; Campanale, A.; Minella, D.; Messina, F.; Butera, O.; Nisii, C.; Mazzarelli, A.; Fontana, C.; Lispi, L.; Maraglino, F.; et al. Epidemiological and Molecular Investigation of the Heater–Cooler Unit (HCU)-Related Outbreak of Invasive Mycobacterium chimaera Infection Occurred in Italy. Microorganisms 2023 , 11 , 2251. [ Google Scholar ] [ CrossRef ]
- Schreiber, P.W.; Kohl, T.A.; Kuster, S.P.; Niemann, S.; Sax, H. The Global Outbreak of Mycobacterium chimaera Infections in Cardiac Surgery—A Systematic Review of Whole-Genome Sequencing Studies and Joint Analysis. Clin. Microbiol. Infect. 2021 , 27 , 1613–1620. [ Google Scholar ] [ CrossRef ]
- Rubinstein, M.; Grossman, R.; Nissan, I.; Schwaber, M.J.; Carmeli, Y.; Kaidar-Shwartz, H.; Dveyrin, Z.; Rorman, E. Mycobacterium intracellulare Subsp. Chimaera from Cardio Surgery Heating-Cooling Units and from Clinical Samples in Israel Are Genetically Unrelated. Pathogens 2021 , 10 , 1392. [ Google Scholar ] [ CrossRef ]
- Mercaldo, R.A.; Marshall, J.E.; Prevots, D.R.; Lipner, E.M.; French, J.P. Detecting Clusters of High Nontuberculous Mycobacteria Infection Risk for Persons with Cystic Fibrosis—An Analysis of U.S. Counties. Tuberculosis 2023 , 138 , 102296. [ Google Scholar ] [ CrossRef ]
- Asadi, T.; Mullin, K.; Roselli, E.; Johnston, D.; Tan, C.D.; Rodriguez, E.R.; Gordon, S. Disseminated Mycobacterium chimaera Infection Associated with Heater-Cooler Units after Aortic Valve Surgery without Endocarditis. J. Thorac. Cardiovasc. Surg. 2018 , 155 , 2369–2374. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Clemente, T.; Spagnuolo, V.; Bottanelli, M.; Ripa, M.; Del Forno, B.; Busnardo, E.; Di Lucca, G.; Castagna, A.; Danise, A. Disseminated Mycobacterium chimaera Infection Favoring the Development of Kaposi’s Sarcoma: A Case Report. Ann. Clin. Microbiol. Antimicrob. 2022 , 21 , 57. [ Google Scholar ] [ CrossRef ]
- Schaeffer, T.; Kuster, S.; Koechlin, L.; Khanna, N.; Eckstein, F.S.; Reuthebuch, O. Long-Term Follow-Up after Mycobacterium chimaera Infection Following Cardiac Surgery: Single-Center Experience. J. Clin. Med. 2023 , 12 , 948. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Trauth, J.; Matt, U.; Kohl, T.A.; Niemann, S.; Herold, S. Blind Spot in Endocarditis Guidelines: Mycobacterium chimaera Prosthetic Valve Endocarditis after Cardiac Surgery—A Case Series. Eur. Heart J. Case Rep. 2023 , 7 , ytad400. [ Google Scholar ] [ CrossRef ]
- Sanavio, M.; Anna, A.; Bolcato, M. Mycobacterium chimaera : Clinical and Medico-Legal Considerations Starting from a Case of Sudden Acoustic Damage. Leg. Med. 2020 , 47 , 101747. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- van Ingen, J. Microbiological Diagnosis of Nontuberculous Mycobacterial Pulmonary Disease. Clin. Chest Med. 2015 , 36 , 43–54. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Wang, H.; Bédard, E.; Prévost, M.; Camper, A.K.; Hill, V.R.; Pruden, A. Methodological Approaches for Monitoring Opportunistic Pathogens in Premise Plumbing: A Review. Water Res. 2017 , 117 , 68–86. [ Google Scholar ] [ CrossRef ]
- Hasse, B.; Hannan, M.M.; Keller, P.M.; Maurer, F.P.; Sommerstein, R.; Mertz, D.; Wagner, D.; Fernández-Hidalgo, N.; Nomura, J.; Manfrin, V.; et al. International Society of Cardiovascular Infectious Diseases Guidelines for the Diagnosis, Treatment and Prevention of Disseminated Mycobacterium chimaera Infection Following Cardiac Surgery with Cardiopulmonary Bypass. J. Hosp. Infect. 2020 , 104 , 214–235. [ Google Scholar ] [ CrossRef ]
- Schreiber, P.W.; Köhler, N.; Cervera, R.; Hasse, B.; Sax, H.; Keller, P.M. Detection Limit of Mycobacterium chimaera in Water Samples for Monitoring Medical Device Safety: Insights from a Pilot Experimental Series. J. Hosp. Infect. 2018 , 99 , 284–289. [ Google Scholar ] [ CrossRef ]
- Daley, C.L.; Iaccarino, J.M.; Lange, C.; Cambau, E.; Wallace, R.J.; Andrejak, C.; Böttger, E.C.; Brozek, J.; Griffith, D.E.; Guglielmetti, L.; et al. Treatment of Nontuberculous Mycobacterial Pulmonary Disease: An Official ATS/ERS/ESCMID/IDSA Clinical Practice Guideline. Clin. Infect. Dis. 2020 , 71 , e1–e36. [ Google Scholar ] [ CrossRef ]
- Lecorche, E.; Haenn, S.; Mougari, F.; Kumanski, S.; Veziris, N.; Benmansour, H.; Raskine, L.; Moulin, L.; Cambau, E.; Aubry, A.; et al. Comparison of Methods Available for Identification of Mycobacterium chimaera . Clin. Microbiol. Infect. 2018 , 24 , 409–413. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Lyamin, A.V.; Ereshchenko, A.A.; Gusyakova, O.A.; Yanchenko, A.V.; Kozlov, A.V.; Khaliulin, A.V. Comparison of Laboratory Methods for Identifying Members of the Family Mycobacteriaceae. Int. J. Mycobacteriol 2023 , 12 , 129–134. [ Google Scholar ] [ CrossRef ]
- Togawa, A.; Chikamatsu, K.; Takaki, A.; Matsumoto, Y.; Yoshimura, M.; Tsuchiya, S.; Nakamura, S.; Mitarai, S. Multiple Mutations of Mycobacterium ntracellulare Subsp. Chimaera Causing False-Negative Reaction to the Transcription-Reverse Transcription Concerted Method for Pathogen Detection. Int. J. Infect. Dis. 2023 , 133 , 14–17. [ Google Scholar ] [ CrossRef ]
- Kuehl, R.; Banderet, F.; Egli, A.; Keller, P.M.; Frei, R.; Döbele, T.; Eckstein, F.; Widmer, A.F. Different Types of Heater-Cooler Units and Their Risk of Transmission of Mycobacterium chimaera during Open-Heart Surgery: Clues from Device Design. Infect. Control Hosp. Epidemiol. 2018 , 39 , 834–840. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Quintás Viqueira, A.; Pérez Romero, C.; Toro Rueda, C.; Sánchez Calles, A.M.; Blázquez González, J.A.; Alejandre Leyva, M. Mycobacterium chimaera in Heater-Cooler Devices: An Experience in a Tertiary Hospital in Spain. New Microbes New Infect. 2021 , 39 , 100757. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Falkinham, J.O., III. Disinfection and Cleaning of Heater-Cooler Units: Suspension- and Biofilm-Killing. J. Hosp. Infect. 2020 , 105 , 552–557. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Hammer-Dedet, F.; Dupont, C.; Evrevin, M.; Jumas-Bilak, E.; Romano-Bertrand, S. Improved Detection of Non-Tuberculous Mycobacteria in Hospital Water Samples. Infect. Dis. Now 2021 , 51 , 488–491. [ Google Scholar ] [ CrossRef ]
- Romano-Bertrand, S.; Evrevin, M.; Dupont, C.; Frapier, J.M.; Sinquet, J.C.; Bousquet, E.; Albat, B.; Jumas-Bilak, E. Persistent Contamination of Heater-Cooler Units for Extracorporeal Circulation Cured by Chlorhexidine-Alcohol in Water Tanks. J. Hosp. Infect. 2018 , 99 , 290–294. [ Google Scholar ] [ CrossRef ]
- Colangelo, N.; Giambuzzi, I.; Moro, M.; Pasqualini, N.; Aina, A.; De Simone, F.; Blasio, A.; Alfieri, O.; Castiglioni, A.; De Bonis, M. Mycobacterium chimaera in Heater–Cooler Units: New Technical Approach for Treatment, Cleaning and Disinfection Protocol. Perfusion 2019 , 34 , 272–276. [ Google Scholar ] [ CrossRef ]
- Ditommaso, S.; Giacomuzzi, M.; Memoli, G.; Garlasco, J.; Curtoni, A.; Iannaccone, M.; Zotti, C.M. Chemical Susceptibility Testing of Non-Tuberculous Mycobacterium strains and Other Aquatic Bacteria: Results of a Study for the Development of a More Sensitive and Simple Method for the Detection of NTM in Environmental Samples. J. Microbiol. Methods 2022 , 193 , 106405. [ Google Scholar ] [ CrossRef ]
- Shrimpton, N.Y.R. Evaluation of Disinfection Processes for Water Heater Devices Used for Extracorporeal Life Support. Perfusion 2019 , 34 , 428–432. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Sarink, M.J.; van Cappellen, W.A.; Tielens, A.G.M.; van Dijk, A.; Bogers, A.J.J.C.; de Steenwinkel, J.E.M.; Vos, M.C.; Severin, J.A.; van Hellemond, J.J. Vermamoeba Vermiformis Resides in Water-Based Heater–Cooler Units and Can Enhance Mycobacterium chimaera Survival after Chlorine Exposure. J. Hosp. Infect. 2023 , 132 , 73–77. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bengtsson, D.; Westerberg, M.; Nielsen, S.; Ridell, M.; Jönsson, B. Mycobacterium chimaera in Heater-Cooler Units Used during Cardiac Surgery–Growth and Decontamination. Infect. Dis. 2018 , 50 , 736–742. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Weitkemper, H.H.; Spilker, A.; Knobl, H.J.; Körfer, R. The Heater-Cooler Unit-A Conceivable Source of Infection. J. Extra Corpor. Technol. 2016 , 48 , 62–66. [ Google Scholar ] [ CrossRef ]
- Foltan, M.; Nikisch, A.; Dembianny, J.; Miano, A.L.; Heinze, J.; Klar, D.; Göbölös, L.; Lehle, K.; Schmid, C. A Solution for Global Hygienic Challenges Regarding the Application of Heater-Cooler Systems in Cardiac Surgery. Perfusion 2023 , 38 , 28–36. [ Google Scholar ] [ CrossRef ]
- Pradal, I.; Esteban, J.; Mediero, A.; García-Coca, M.; Aguilera-Correa, J.J. Contact Effect of a Methylobacterium sp. Extract on Biofilm of a Mycobacterium chimaera Strain Isolated from a 3T Heater-Cooler System. Antibiotics 2020 , 9 , 474. [ Google Scholar ] [ CrossRef ]
- Masaka, E.; Reed, S.; Davidson, M.; Oosthuizen, J. Opportunistic Premise Plumbing Pathogens. A Potential Health Risk in Water Mist Systems Used as a Cooling Intervention. Pathogens 2021 , 10 , 462. [ Google Scholar ] [ CrossRef ]
- Treglia, M.; Pallocci, M.; Ricciardi Tenore, G.; Castellani, P.; Pizzuti, F.; Bianco, G.; Passalacqua, P.; De Luca, L.; Zanovello, C.; Mazzuca, D.; et al. Legionella and Air Transport: A Study of Environmental Contamination. Int. J. Environ. Res. Public Health 2022 , 19 , 8069. [ Google Scholar ] [ CrossRef ]
- Glassmeyer, S.T.; Burns, E.E.; Focazio, M.J.; Furlong, E.T.; Gribble, M.O.; Jahne, M.A.; Keely, S.P.; Kennicutt, A.R.; Kolpin, D.W.; Medlock Kakaley, E.K.; et al. Water, Water Everywhere, but Every Drop Unique: Challenges in the Science to Understand the Role of Contaminants of Emerging Concern in the Management of Drinking Water Supplies. Geohealth 2023 , 7 , e2022GH000716. [ Google Scholar ] [ CrossRef ]
- Ortiz-Martínez, Y. Mycobacterium chimaera : An under-Diagnosed Pathogen in Developing Countries? J. Hosp. Infect. 2017 , 97 , 125–126. [ Google Scholar ] [ CrossRef ]
- Becker, J.B.; Moisés, V.A.; Guerra-Martín, M.D.; Barbosa, D.A. Epidemiological Differences, Clinical Aspects, and Short-Term Prognosis of Patients with Healthcare-Associated and Community-Acquired Infective Endocarditis. Infect. Prev. Pract. 2024 , 6 , 100343. [ Google Scholar ] [ CrossRef ]
- Ferrara, S.D.; Baccino, E.; Bajanowski, T.; Boscolo-Berto, R.; Castellano, M.; De Angel, R.; Pauliukevičius, A.; Ricci, P.; Vanezis, P.; Vieira, D.N.; et al. Malpractice and Medical Liability. European Guidelines on Methods of Ascertainment and Criteria of Evaluation. Int. J. Leg. Med. 2013 , 127 , 545–557. [ Google Scholar ] [ CrossRef ]
Click here to enlarge figure
| References | Author, Year | N. of Patients | Surgery | Mean Time of Presentation If Previous Surgery | Setting (Country) | Organ and/or Tissue Involved |
|---|---|---|---|---|---|---|
| [ ] | (Bills et al., 2009) | 1 | None | Na | Not healthcare (USA) | Lung, nodules in chronic obstructive pulmonary disease |
| [ ] | (Cohen-Bacrie et al., 2011) | 1 | None | Na | Possible frequent healthcare contact (Réunion Island, FR) | Lung infections in cystic fibrosis |
| [ ] | (Alhanna et al., 2012) | 1 | None | Na | Not healthcare (Germany) | Lung infection |
| [ ] | (Gunaydin et al., 2013) | 5 (of 90) | None | Na | Possible healthcare contact (Turkey) | Lung (reassessment of sputum specimens) |
| [ ] | (Boyle et al., 2015) | 125 (of 448) | None | Na | Possible healthcare contact (USA) | Lung (reassessment of sputum specimens) |
| [ ] | (Mwikuma et al., 2015) | 1 (of 54) | None | Na | Not healthcare (Zambia) | Lung (reassessment of sputum specimens) |
| [ ] | (Moon et al., 2016) | 11 | None | Na | Not healthcare (South Korea) | Lung infection (reassessment of sputum specimens) |
| [ ] | (Moutsoglou et al., 2017) | 1 | None | Na | Not healthcare (USA) | Disseminated with spinal osteomyelitis and discitis |
| [ ] | (Bursle et al., 2017) | 1 | Tricuspid valve repair and mitral annuloplasty | 13 months | Underwent surgery (Australia) | Disseminated |
| [ ] | Kim et al., 2017 | 8 (of 91) | None | Na | Possible healthcare contact (Korea) | Lung (reassessment of sputum specimens) |
| [ ] | (Chand et al., 2017) * | 4 | Valvular cardiac surgery | 1.15 (0.25–5.1) years | Underwent surgery (UK) | 1 osteomyelitis and 3 disseminated |
| [ ] | (Truden et al., 2018) | 49 (of 102) | None | Na | Possible healthcare contact (Slovenia) | Lung (reassessment of sputum specimens) |
| [ ] | (Larcher et al., 2019) | 4 | None | Na | Possible frequent healthcare contact (France) | Lung (reassessment of sputum specimens in cystic fibrosis) |
| [ ] | (Shafizadeh et al., 2019) * | 5 | Valvular cardiac surgery | 20.6 (14–29) months | Underwent surgery (USA) | Disseminated with liver infection |
| [ ] | (Rosero and Shams, 2019) | 1 | None but operating room nurse 10 years ago | >10 years | Possible frequent healthcare contact (USA) | Lung infection |
| [ ] | (Watanabe et al., 2020) | 1 | None | Na | Not healthcare (Japan) | Tendons, hand tenosynovitis |
| [ ] | (Chen et al., 2020) | 28 | None | Na | Not healthcare (Taiwan) | Lung infection (reassessment of sputum specimens) |
| [ ] | (Maalouly et al., 2020) | 1 | Kidney transplantation | One week | Underwent surgery (Belgium) | Kidney, urinary tract infection in a kidney transplant recipient with concomitant Mycobacterium malmoense lung infection and fibro anthracosis |
| [ ] | (de Melo Carvalho et al., 2020) | 1 | None | Na | Possible healthcare contact (Portugal) | Disseminated in B-cell lymphoma |
| [ ] | (Sharma et al., 2020) | 2 | None | Na | Not healthcare (India) | Meninges, meningitis |
| [ ] | (Zabost et al., 2021) | 88 (of 200) | None | Na | Possible healthcare contact (Poland) | Lung infection (reassessment of sputum specimens) |
| [ ] | (Kim et al., 2021) | 4 (of 320) | None | Na | Possible healthcare contact (Korea) | Lung infection (reassessment of sputum specimens) |
| [ ] | (Kavvalou et al., 2022) | 1 | None | Na | Possible healthcare contact (Germany) | Central venous catheter infection in cystic fibrosis |
| [ ] | (Robinson et al., 2022) | 1 | None | Na | Not healthcare (USA) | Lung infection in drug abuser |
| [ ] | (Ahmad et al., 2022) | 1 | None | Na | Not healthcare (USA) | Lung infection in sarcoidosis |
| [ ] | (George et al., 2022) | 1 | None | Na | Not healthcare (India) | Skin, periapical abscess with chin ulcer |
| [ ] | (Lin et al., 2022) | 1 | None | Na | Possible frequent healthcare contact (Taiwan) | Disseminated in adult-onset immunodeficiency syndrome |
| [ ] | (Łyżwa et al., 2022) | 1 | None | Na | Not healthcare (Poland) | Lung infection in silicosis |
| [ ] | (McLaughlin et al., 2022) | 1 | Coronary artery bypass grafting | 1 year | Underwent surgery (USA) | Tendons, hand tenosynovitis in ipsilateral elbow wound in fisherman |
| [ ] | (Gross et al., 2023) | 23 | None | Na | Healthcare (USA) | Lung infections in cystic fibrosis (genomic analysis for cluster correlation to hospital outbreaks) |
| [ ] | (Azzarà et al., 2023) | 1 | None | Na | Possible healthcare contact (Italy) | Lung infection in lung adenocarcinoma treated with immune checkpoint inhibitors |
| [ ] | (Pradhan et al., 2023) | 1 | Bioprosthetic mitral valve replacement | 7 years | Underwent surgery (Australia) | Spinal osteomyelitis and discitis |
| [ ] | (Garcia-Prieto et al., 2024) | 1 | None | Na | Not healthcare (Spain) | Lung infection in fibro anthracosis |
| [ ] | (Paul et al., 2024) | 1 | None | Na | Possible healthcare contact (UK) | Lung infection in unilateral pulmonary artery agenesis on the right side |
| The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
Share and Cite
Bolcato, V.; Bassetti, M.; Basile, G.; Bianco Prevot, L.; Speziale, G.; Tremoli, E.; Maffessanti, F.; Tronconi, L.P. The State-of-the-Art of Mycobacterium chimaera Infections and the Causal Link with Health Settings: A Systematic Review. Healthcare 2024 , 12 , 1788. https://doi.org/10.3390/healthcare12171788
Bolcato V, Bassetti M, Basile G, Bianco Prevot L, Speziale G, Tremoli E, Maffessanti F, Tronconi LP. The State-of-the-Art of Mycobacterium chimaera Infections and the Causal Link with Health Settings: A Systematic Review. Healthcare . 2024; 12(17):1788. https://doi.org/10.3390/healthcare12171788
Bolcato, Vittorio, Matteo Bassetti, Giuseppe Basile, Luca Bianco Prevot, Giuseppe Speziale, Elena Tremoli, Francesco Maffessanti, and Livio Pietro Tronconi. 2024. "The State-of-the-Art of Mycobacterium chimaera Infections and the Causal Link with Health Settings: A Systematic Review" Healthcare 12, no. 17: 1788. https://doi.org/10.3390/healthcare12171788
Article Metrics
Article access statistics, supplementary material.
ZIP-Document (ZIP, 83 KiB)
Further Information
Mdpi initiatives, follow mdpi.

Subscribe to receive issue release notifications and newsletters from MDPI journals
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- HHS Author Manuscripts

Mouse models for infectious diseases caused by Staphylococcus aureus
Staphylococcus aureus - a commensal of the human skin, nares and gastrointestinal tract - is also a leading cause of bacterial skin and soft tissue infection (SSTIs), bacteremia, sepsis, peritonitis, pneumonia and endocarditis. Antibiotic-resistant strains, designaed MRSA (methicillin-resistant S. aureus ), are common and represent a therapeutic challenge. Current research and development efforts seek to address the challenge of MRSA infections through vaccines and immune therapeutics. Mice have been used as experimental models for S. aureus SSTI, bacteremia, sepsis, peritonitis and endocarditis. This work led to the identification of key virulence factors, candidate vaccine antigens or immune-therapeutics that still require human clinical testing to establish efficacy. Past failures of human clinical trials raised skepticism whether the mouse is an appropriate model for S. aureus disease in humans. S. aureus causes chronic-persistent infections that, even with antibiotic or surgical intervention, reoccur in humans and in mice. Determinants of S. aureus evasion from human innate and adaptive immune responses have been identified, however only some of these are relevant in mice. Future research must integrate these insights and refine the experimental mouse models for specific S. aureus diseases to accurately predict the failure or success for candidate vaccines and immune-therapeutics.
Staphylococcus aureus
Staphylococcus aureus is a Gram-positive bacterium that colonizes the skin, nares and gastrointestinal tract of humans ( 1 ). Approximately 20% of the human population are stably colonized while 30% are colonized in a variable manner ( 1 , 2 ). S. aureus is also a pathogen that causes invasive disease, predominantly skin and soft tissue infections (SSTI) but also bacteremia, sepsis, pneumonia, osteomyelitis and endocarditis ( 1 ). The pathological hallmark of S. aureus infection is the formation of purulent abscess lesions that are formed around a nidus of the pathogen, primarily via the infiltration of neutrophils ( 3 ). In humans, S. aureus infection does not lead to the development of protective immune responses and chronic persistent or recurrent infections are common ( 4 ). Some isolates of S. aureus cause toxic-shock syndrome, exfoliative skin disease, and enteritis in humans ( 1 ). Secreted toxins are the key virulence determinants for these diseases ( 5 , 6 ) and transfer of the corresponding genes among staphylococcal strains involves specific bacteriophages ( 7 ).
S. aureus is also an important pathogen of live-stock, causing large scale infections in ruminants (sheep, goats, cows), poultry and pigs ( 8 ). Molecular epidemiological data suggest that a common pathogenic S. aureus clone associated with ruminants originated in humans ( 9 ). This strain adapted to its chosen niche more than 11,000 years ago, at a time when farming domesticated animals became common practice, and then diversified ( 9 ). Similar jumps from humans to new hosts occurred for several different clinical lineages ( 10 ). Adaptation to new hosts required a combination of gene loss, allelic diversification, and acquisition of mobile genetic elements, for example elements that support the expression of unique von-Willebrand factor binding protein alleles in S. aureus strains that infect ruminants and horses ( 11 ). Nevertheless, the core genome of ruminant associated S. aureus is stable and can lead to reciprocal transmission of newly emerging clones into the human population ( 12 ). This type of pathogen introduction occurs on a global scale and is associated with transport of live-stock or people ( 13 ). It has led to outbreaks of human S. aureus disease in countries that otherwise have low prevalence for staphylococcal disease ( 9 ).
The genome of S. aureus strains varies in size (2.6-2.9 MB), based on the presence of prophages and pathogenicity islands ( 14 , 15 ). Nevertheless, all S. aureus isolates encompasses a core genome for the functional expression of genes that are shared by most if not all clinically relevant strains ( 16 ). For example, S. aureus isolates generally coagulate blood, agglutinate in plasma, and bind to animal immunoglobulin ( 17 ). Although nasal colonization is asymptomatic for most individuals, it represents a risk factor for hospital-acquired infection ( 18 ). Other risk factors for nosocomial infection include indwelling catheters, endotracheal intubation, medical implants, trauma, diabetes, immunosuppression and immunosuppressive therapy, hemodialysis and peritoneal dialysis ( 19 , 20 ). In the United States, S. aureus is the single most frequent cause of hospital-acquired infectious disease mortality ( 21 ).
Massive use of antibiotics in animals and humans led to the selection of drug-resistant strains, designated MRSA for methicillin-resistant S. aureus ( 22 ). Broad spectrum β-lactam resistance is caused by MecA, a penicillin-binding protein that cannot be inhibited by β-lactamase resistant β-lactam or cephalosporin compounds ( 22 ). Recommended therapeutics against MRSA strains are vancomycin, daptomycin and linezolid ( 23 - 25 ). However, vancomycin-resistant (VRSA) strains, which acquired genes for the synthesis of altered peptidoglycan precursors from enterococci, have been isolated ( 26 - 28 ). S. aureus strains have also evolved resistance mechanisms to daptomycin and linezolid shortly after these compounds were approved for clinical use ( 28 - 30 ). MRSA infections occur frequently in American hospitals and in the community ( 21 ). The public health crisis of community- and hospital-acquired MRSA infections may be addressed through the development of vaccines, however all clinical trials that sought to address this problem have thus far failed ( 31 ). Healthcare providers currently have limited treatment options, a situation reminiscent of the pre-antibiotic era ( 32 ). The National Academy of Science’s Institute of Medicine (IOM) placed comparative effectiveness research on MRSA as one of the top 25 priorities for national investment. S. aureus was the only bacterial pathogen addressed in the IOM report.
Staphylococcus aureus infections of mice
When designed to recapitulate human disease, animal studies with infectious agents aim to provide experimental proof for the molecular basis of pathogenesis, the establishment of protective immunity and the molecular mechanisms whereby immunity is achieved ( 33 - 35 ). Over the past forty years, infectious diseases research championed the mouse as a model for human infectious diseases, generating a plethora of reagents that enable rapid advances. Another important factor favoring the mouse is costs. Small body size, short gestation and large litter size render mice less expensive to rear than any other mammal currently used for infectious diseases research ( 35 - 37 ).
Mice are, however, not a natural host for human clinical S. aureus isolates and require genetic adaptation to cause transmissible disease in mouse facilities ( 38 ). Key clinical presentation of natural S. aureus infection is the development of preputial gland abscess lesions in male mice as well as nasal and gastrointestinal colonization ( 38 ). Because most experimental approaches are designed towards elucidating the virulence factors of human clinical isolates and deriving protective vaccines for humans, mouse isolates have not yet been examined in detail. Nevertheless, mice have served as the premier experimental model to study S. aureus blood-borne persistent and metastatic abscess formation ( 3 ), septic arthritis ( 39 ), sepsis ( 40 ), neonatal sepsis and meningitis ( 41 ), endocarditis ( 42 ), peritonitis ( 43 ), subcutaneous skin infection ( 44 ) and pneumonia ( 45 ). This work provided several key insights. First, S. aureus can induce a diverse spectrum of diseases in mice - similar to what is observed in humans - albeit that the infectious dose required to produce a desired pathology varies widely. Second, the genetic determinants of S. aureus required to produce specific pathologies vary between the different disease models but are generally consistent when analyzing different staphylococcal strains ( 3 , 45 ). Third, some models require a very large inoculum to elicit disease pathology, and one cannot expect to identify many of the key genetic determinants of S. aureus that would support pathogenesis in humans, where a lower inoculum may suffice ( 46 ). Fourth, the pathogenesis of different S. aureus infectious diseases appears to require multiple virulence factors and these can differ between disease types ( 17 , 47 , 48 ). Fifth, some secreted factors play key roles in neutralizing human innate defenses and are expressed by virtually all human clinical isolates, yet these factors are not adapted to mice and play no role in the various disease models ( 49 , 50 ). Sixth, both immune-competent inbred and outbred mice can be used as disease models although the infectious dose required to produce disease can vary by a factor of 10-100 ( 51 ).
Below we discuss the most frequently used mouse models for S. aureus disease, describe the key virulence factors, and the relevance of each model for human disease. We point to important mechanisms for pathogen survival, and describe how each model was used for the discovery of protective antigens. It has been argued that mouse models cannot accurately identify the protective antigens of S. aureus and that mouse studies have misinformed the development of vaccine antigens ( 52 ). This criticism is not entirely justified. Past vaccine failures can also be explained by mismatches between disease model and trial design, by unexpected safety issues and by ignoring the immune evasive attributes of S. aureus ( 53 ).
Preparing S. aureus for experimental infection
Blood-borne spread of staphylococci can generate abscess lesion in any human organ tissue. Initially, S. aureus strains were isolated from specific organ tissues with the notion that unique genetic determinants or the variation of certain virulence genes may favor replication at these sites ( 54 ). For example, strains isolated from infected endocardium or infected joints were used to examine infective endocarditis and arthritis in animals, respectively ( 54 ). Extensive DNA sequence analysis of clinical isolates has demonstrated that S. aureus strains, although endowed with variable genetic traits ( 55 ), invariably retain the ability for blood-borne dissemination and for the seeding of abscess lesions ( 15 , 16 ). Such insight enables the field to focus on a few well characterized strains, for example the methicillin-sensitive isolate S. aureus Newman ( 55 ) and the CA-MRSA isolate USA300 (LAC) ( 56 ), to conduct comparative analyses of virulence factors and vaccine antigens. To standardize the inoculation of staphylococci into animals, we recommend that bacteria are grown as overnight cultures in tryptic soy broth with rotation at 37°C, diluted 1:100 into fresh broth and incubated further to absorbance at 600 nm (A 600 ) 0.5. Staphylococci in culture aliquots should be sedimented by centrifugation, bacteria washed and suspended in sterile PBS to the desired volume and inoculum. Each inoculum should be quantified by spreading aliquots on agar plates and enumerating colony forming units (CFU).
Skin infection
Ogston first isolated S. aureus from the pus of surgical wound infections and showed that, when injected into the subcutaneous tissue of experimental animals, this material could elicit abscess lesions in guinea pigs and mice ( 57 , 58 ). Infections in humans may occur at sites where the skin barrier function has been breached (wound or surgical site infections) or without apparent breaches, for example at hair follicles (folliculitis), as bullous or superficial lesions (impetigo), deep (furuncles) or confluent abscesses (carbuncles) ( 1 ). S. aureus infections of the human skin elicit purulent exudate, which drains from the infectious site. To induce skin infections in mice, the hair on the back or flank of mice is shaved and a suspension of 10 7 -10 9 CFU S. aureus in PBS is injected into subcutaneous tissues ( 44 , 59 ). Within 24 hours, bacteria elicit inflammatory responses and cause an indurated swelling, which increases over 5-7 days to a size of 30-100 mm 2 ( 59 ). Abscess lesions are gradually resolved over the next 7-9 days either with or without drainage of purulent material ( 44 , 46 ). Depending on the S. aureus strain tested and its production of secreted α-hemolysin (α-toxin), subcutaneous skin lesions are associated with superficial dermonecrosis, which heals at a similar rate as the resolution of subcutaneous abscess lesions ( 46 ).
S. aureus strains require the structural gene for α-hemolysin ( hla ) to produce wild-type size lesions and dermonecrosis in the mouse model for skin disease ( 46 ). Immunization of mice with Hla H35L , a non-toxigenic variant ( 60 ), elicits neutralizing antibodies that provide protection against Hla-mediated pathological effects but cannot prevent the establishment of smaller sized lesions without dermonecrosis ( 46 ). Similar protective effects are achieved in passive transfer experiments with monoclonal antibodies that neutralize Hla ( 61 ). ADAM10 (a disintegrin and metalloproteinase domain-containing protein 10) serves as the receptor for Hla and is conserved in humans and mice ( 62 ). The Hla-ADAM10 interaction leads to disruption of host cell membrane junctions, a process that exacerbates necrosis of the epidermis and dermis ( 46 , 62 , 63 ).
MRSA strains of the USA300 lineage are causing an epidemic of community-acquired skin infections in the United States ( 64 , 65 ). These strains express a bacteriophage encoded leukocidin, the pore-forming cytotoxin PVL ( 66 , 67 ), which binds to the complement receptors C5aR and C5L2 of human leukocytes but not to the complement receptors of murine immune cells ( 68 ). Not surprisingly, deletion of the structural genes for PVL does not affect the pathogenesis of USA300 variants in the mouse skin infection model ( 69 , 70 ).
Phenol soluble modulins (PSMs) contribute to the pathogenesis of mouse skin infections. Originally identified in S. epidermidis ( 71 ), subsequent studies in MRSA described two types of PSMs: α-PSMs are relatively short (20-25 amino acid long) and form an amphipathic α-helix, whereas β-PSMs are longer ( ~ 50 amino acid long) and only their C-terminal domain may adopt an α-helical structure ( 72 , 73 ). PSMs are not synthesized as signal-peptide bearing precursors and require a dedicated ATP-binding cassette (ABC) transporter for secretion as formylated peptides ( 74 ). PSMs elicit proinflammatory responses by interacting with the formyl peptide receptor 2 (FPR2) of host cells including neutrophils ( 74 , 75 ). In particular, PSMα3 has been proposed to activate, attract and lyse neutrophils ( 72 ). S. aureus continues to produce PSMs following its uptake by neutrophils, suggesting that PSMs may perhaps promote an intracellular lytic activity ( 76 ). In agreement with these hypotheses, S. aureus variants unable to produce α-PSMs are associated with reduced dermonecrosis ( 72 , 74 ).
The mouse model for S. aureus skin infection has been modified to examine the contribution of specific skin cells or tissue structures and of specific immune cells such as neutrophils or macrophages. Some protocols damage the skin (removal of superficial keratinocytes, incision or heat damage) prior to inoculation of S. aureus, whereas other protocols inoculate staphylococci along with implanted foreign material (sutures, dextran beads, cotton dust) ( 77 - 80 ). These modifications allow investigators to reduce the staphylococcal challenge dose required to cause pathologic lesions.
Depletion of immune cells via specific monoclonal antibodies (anti-Ly6G) or chemotherapy (cyclophosphamide) causes a dramatic reduction in the challenge dose required for skin infection ( 81 , 82 ). In contrast to immune-competent mice, leukopenic mice cannot contain staphylococci in the skin and develop systemic infections with rapidly lethal outcome ( 83 ). Similar observations have been made in humans, where iatrogenic leukopenia or hereditary defects in the NADPH oxidase or in the respiratory burst of myeloid cells are associated with increased susceptibility towards S. aureus infection ( 84 ). Humans with abnormal T cell function (including HIV/AIDS patients), atopic dermatitis, and hyper-IgE syndrome, also display increased susceptibility toward S. aureus cutaneous infection ( 85 - 88 ). These phenotypes can be recapitulated in mice with similar skin conditions or immune defects ( 80 ). For example skin lesions of S. aureus -infected MyD88- and IL-1R-deficient mice, and to a lesser extend TLR2-deficient mice, are substantially larger in size, with higher staphylococcal load, as compared to lesions in wild-type mice ( 89 ). The severity of skin infections correlates with the decreased recruitment of neutrophils to the site of infection ( 89 ). Similarly, mice deficient in dendritic epidermal T cells (γδ T cells) and IL-17R develop larger skin lesions with higher bacterial load and impaired neutrophil recruitment ( 90 ). IL-17 producing T lymphocytes (T H 17) play an important role in controlling S. aureus cutaneous infection by recruiting neutrophils to the infectious site ( 91 ). A growing body of evidence implicates T H 17 cells as critical for protection against skin and lung infections, whereas T H 17/IL-17 responses may be less critical in other tissues ( 92 , 93 ).
Thus, immune-competent mice (C57BL/6 or BALB/c) represent a good model for the study of S. aureus skin infections and the characterization of staphylococcal virulence factors. The subcutaneous injection of a relatively large inoculum prevents the identification of virulence factors acting at early stages of disease pathogenesis (adherence to skin structures, invasion across epidermal layers). As some secreted factors of S. aureus require human specific interactions, the mouse model can likely be improved by generating transgenic animals expressing specific human genes or by developing mouse models in which the mouse innate or adaptive immune system is replaced by its human counterpart ( 94 ). For example, expression of human complement receptors C5aR in murine myeloid cells could restore susceptibility toward staphylococcal PVL. Approximately one-fifth of SSTI patients receiving antibiotic therapy develop recurrent skin infections with the same strain ( 95 ). It will be important to determine whether recurrent skin infection occurs in mice and, if so, what genetic determinants of staphylococci promote this disease process.
Bacteremia and metastatic abscess formation
Invasive staphylococcal disease in humans is associated with staphylococcal bacteremia and the formation of abscess lesions in many different organ tissues ( 96 ). In mice, intravenous inoculation of S. aureus also triggers dissemination of blood-borne bacteria into organ tissues, where they establish abscess lesions in skeletomuscular, vascular, brain, lung, heart, liver, spleen and kidney tissues ( 3 ). Briefly, mice are injected into the tail vein or the periorbital venous plexus with 10 6 -10 7 CFU, which causes staphylococcal bacteremia ( 3 ). The bacteria eventually exit from the blood stream but can be recovered from peripheral organs, for example renal tissues, where they attract large immune cell infiltrates. Hematoxylin-eosin staining of thin-sectioned tissue samples provides the first histopathologic evidence for S. aureus infection, as early as 48 hours following intravenous inoculation ( 3 ). By day 4, renal tissues harbor multiple abscess lesions that, on histopathologic examination are characterized by a bacterial nidus ( staphylococcal abscess community ) encapsulated by eosinophilic fibrin deposits. These fibrin deposits shield staphylococci from surrounding immune cell infiltrates. During early stages of abscess development (days 2-5), immune cells infiltrates are comprised predominantly of neutrophils, arranged in concentric layers of apoptotic, healthy and apoptotic PMNs ( 3 ). Macrophages and lymphocytes are positioned in the periphery of these lesions ( 97 ). By day 15 following inoculation, T and B lymphocytes as well as macrophages are recruited into abscess lesions, which slowly progress towards the organ surface and eventually rupture to release purulent material with staphylococci into circulation ( 3 ). Released staphylococci seed new abscesses at other sites and, by day 30-50, accumulating lesions eventually promote lethality ( 3 ).
S. aureus Newman variants have been analyzed for their phenotypic defects in the renal abscess model. Sortase A mutants, defective for anchoring any one of 19 surface proteins with LPXTG sorting signals to the cell wall envelope, are unable to cause abscess lesions ( 3 ). In contrast, mutations in individual surface protein genes cause either no significant defect or display a reduction in the number and size of abscess lesions and in staphylococcal load ( 3 ). Mutants that display the most severe defects harbor mutations in genes for the following surface proteins: staphylococcal protein A (SpA), iron-regulated surface determinant A (IsdA) and B (IsdB), adenosine synthase A (AdsA), as well as serine-aspartate repeat protein D (SdrD) ( 3 ). Both IsdA and IsdB promote heme-iron scavenging during infection ( 98 , 99 ) and antibodies against these proteins provide partial protection by reducing the number of abscess lesions and the bacterial load in renal tissues ( 100 - 102 ). The IsdB vaccine was subjected to a clinical trial in humans to reduce post-surgical mediastinal and superficial wound infections as well as bacteremia in patients undergoing cardiothoracic surgery ( 103 ). The trial was ended because multiorgan dysfunction and mortality following S. aureus infection occurred more frequently in individuals receiving the IsdB vaccine than in control cohorts and IsdB immunization did not show clinical benefit ( 103 ). Antibodies against SdrD also provide partial protection against S. aureus renal abscess formation, albeit that the molecular basis for protection is not known since the exact function of SdrD is still unknown ( 101 ). A similar protective effect is achieved when the CnaB domain of SdrE, which is conserved among members of this protein family (SdrD and SdrE) ( 104 ), is used as a vaccine antigen ( 105 ).
SpA binds to human and animal immunoglobulins (IgG, IgA, IgD, IgE and IgM) either at Fcγ, the C H 2-C H 3 hinge region of IgG involved in binding to complement factor C1q and Fcγ-receptors, or the heavy chain variable region (Fab) of V H 3 class antibodies ( 106 - 108 ). The former activity neutralizes antibody effector functions for opsonophagocytosis ( 109 ), whereas the latter activates V H 3-clan B cell receptors (IgM), resulting in non-productive B cell expansions and suppression of adaptive immune responses ( 110 , 111 ). SpA-neutralizing polyclonal or monoclonal antibodies provide partial protection in the renal abscess model but also neutralize the B cell superantigen activity of staphylococci to promote antibody responses against almost all secreted products of S. aureus ( 112 , 113 ). When used as a vaccine antigen, SpA KKAA , a non-toxigenic variant that cannot bind immunoglobulin, elicits neutralizing antibodies and partial protection against S. aureus renal abscess disease ( 112 ). Further, SpA KKAA immunization enhances antibody responses against many secreted products of S. aureus in animals that are exposed to this pathogen ( 112 ). SpA-mediated B cell superantigen activity is thought to suppress pathogen-specific antibody responses and the development of protective immunity ( 114 ). In agreement with this, infection of mice with spa variants that lack B cell superantigen activity elicits broad-spectrum antibody responses and promotes protective immunity ( 111 ).
AdsA, another surface protein requiring sortase A for envelope assembly, catalyzes the conversion of ATP, ADP and AMP to adenosine, a signal for mammalian immune suppression ( 115 , 116 ). AdsA also converts 5’ and 3’ monophosphate cleavage products of DNA, derived from staphylococcal nuclease activity with NETs (neutrophil extracellular DNA traps) ( 117 ), into deoxyadenosine, thereby inducing apoptosis in macrophages ( 97 ). This mechanism supports abscess formation and staphylococcal escape from phagocytic clearance in these lesions, which for several days remain devoid of macrophages ( 97 ).
S. aureus secretes two polypeptides, coagulase (Coa) and von-Willebrand Factor binding protein (vWbp), that associate with and activate prothrombin to generate staphylothrombin complexes, thereby cleaving fibrinogen and generating fibrin clots ( 118 , 119 ). This activity, which can be monitored in vitro with the coagulase test, is essential for the establishment of renal abscesses, as coa/vwb mutants are unable to generate these lesions ( 119 ). When used as vaccine antigens, Coa and vWbp raise neutralizing antibodies that protect against abscess lesions ( 119 ). Of note, α-hemolysin does not play a key role in the establishment of abscess lesions ( 3 ). Capsular polysaccharides (CP5/CP8) and poly- N -acetylglucosamine (PNAG) have been advocated as important vaccine antigens, raising antibody responses protective against staphylococcal abscess formation in renal tissues ( 120 - 123 ). Compared with other virulence determinants, the traits for capsule or PNAG production are not critically important for disease pathogenesis in S. aureus Newman and in the CA-MRSA isolate USA300 LAC, presumably because encapsulation is either diminished or absent ( 3 , 124 ).
In summary, staphylococcal abscess formation can be thought of as a developmental process that occurs in four discrete stages ( 17 ). Following intravenous inoculation, S. aureus requires specific genes to survive in the blood stream of infected mice for subsequent deposition into peripheral organ tissues (Stage I). Of note, staphylococci can disseminate as extracellular pathogens but also when located within neutrophils ( 125 ). Staphylococcal invasion of host tissues triggers massive immune cell infiltrations with neutrophils predominating early during infection (Stage II). Due to the formation of a fibrin capsule and other virulence strategies, staphylococci replicate as abscess communities and manipulate immune cell infiltrates to promote tissue destruction and abscess maturation at organ surfaces (Stage III). Liquefaction necrosis and drainage of pus releases staphylococci for renewed dissemination to new sites within the infected host (Stage IV).
Blood stream infections are a risk factor for septic arthritis in humans and the metastatic abscess model can also be used to study S. aureus septic arthritis in mice ( 54 ). Following inoculation of staphylococci, mice are weighed daily and examined for arthritis and clinical appearance. At timed intervals, one pair of limbs (fore and hind) is used for histopathologic examination of joints (synovitis), while paws are examined for the bacterial load. Alternatively, bacteria may be injected directly in the knee joint ( 126 ). In this model, 3.6×10 4 CFU of strain Newman in 20 μl of PBS are injected into the joint and animals are killed 3 days after inoculation for histopathologic examination of synovitis. Synovitis is defined as synovial membrane thickness of over two cell layers and infiltration of inflammatory cells to the extra-articular space with cartilage or bone destruction ( 126 ). A histopathology index scores the severity of synovitis and tissue destruction ( 126 ). S. aureus sortase A, protein A and clumping factor A ( vide infra ) are virulence factors in this model, whereas Hla and IsdA/IsdB are not ( 127 - 131 ).
S. aureus is a common cause of sepsis, a frequently fatal, systemic inflammation with multiple organ dysfunction, typically triggered by immune responses towards bacterial replication in blood ( 132 ). Intravenous inoculation of 5×10 7 -5×10 8 CFU S. aureus causes infected animals to develop septic shock with lethal outcome within 12-48 hours ( 119 ). Animals present with clinical signs of disease within 2-3 hours, including ruffled fur, diminished activity and appetite, hunched posture, loss of movement, diarrhea, dehydration and labored breathing ( 3 ). During necropsy, staphylococci can be isolated from blood and from all organ tissues examined. S. aureus Newman variants lacking sortase A are avirulent in this model system and do not precipitate a lethal outcome over 14 days following blood stream inoculation ( 133 ). Two surface proteins play a key role in sepsis pathogenesis, AdsA ( vide supra ) and ClfA ( 133 ). Clumping factor A (ClfA) binds to the D domain of fibrinogen and fibrin ( 134 - 136 ). When expressed in human or animal plasma together with Coa and vWbp, ClfA mediates staphylococcal association with fibrin filaments, a phenotype that has been designated clumping or agglutination ( 133 ). ClfA association with fibrin filaments protects staphylococci from macrophages ( 137 ) and also promotes vascular occlusions by agglutinated bacteria in organ tissues ( 133 ). A triple mutant strain lacking coa, vwb and clfA is avirulent in the mouse sepsis model, whereas clfA or coa/vwb variants are severely attenuated ( 119 , 133 ). The pathogenesis of S. aureus sepsis can be perturbed with antibodies that neutralize the biochemical activities of Coa, vWbp and ClfA or with direct thrombin inhibitors, small molecules that also block staphylothrombin cleavage of fibrinogen ( 119 , 133 ). ClfA, Coa and vWbp-derived vaccines can prevent S. aureus sepsis in mice ( 138 ) and mice with a genetic defect in fibrinogen, which abolishes its association with ClfA, are less susceptible to S. aureus induced sepsis than wild-type mice ( 139 ).
S. aureus hla mutants display delayed time-to-death and increased survival in the mouse sepsis model, likely because ADAM10 activation by the toxin is a key factor for the increased vascular permeability associated with disease ( 140 ). S. aureus secretes several other pore forming toxins, LukED and LukAB, that cause injury to myeloid cells by interacting with their G-protein coupled receptor molecules ( 141 - 144 ). The severity of S. aureus induced sepsis can be ameliorated by depletion of neutrophils that are targeted by LukED in a manner requiring specific chemokine receptors ( 141 , 145 ). LukAB association with its CD11b receptor occurs in a host specific manner, as this toxin is capable of injuring human neutrophils but not mouse neutrophils ( 144 ). The possibility of targeting leukocidins (LukED, LukAB, PVL) as vaccine antigens represents an exciting frontier that awaits the development of refined mouse models with functional human chemokine receptors on myeloid cells.
Peritonitis
S. aureus is a frequent cause of peritonitis in individuals with end-stage renal disease and continuous ambulatory peritoneal dialysis ( 146 ). When injected with staphylococci into the peritoneal cavity, mice require a very large inoculum to develop lethal disease: LD 50 =5×10 8 CFU and LD 90 = 6×10 9 CFU for S. aureus Newman ( 43 ). The animals typically succumb within 12-24 hours of challenge and survivors of lower challenge doses harbor intraperitoneal abscess lesions filled with large numbers of immune cells, mostly neutrophils and macrophages, and surrounded by layers of fibrin and collagen ( 43 ). Very few staphylococci enter the blood stream and the establishment of metastatic abscess lesions is rare. On the other hand, lesions that were formed within the recesses of the peritoneal cavity are often associated with tissues of organs that are covered with peritoneal lining, e.g. the kidneys. Following necropsy, staphylococci are found in homogenized renal tissues, although the bacteria were located in the peritoneal cavity but not in renal parenchyma ( 43 ). The ease of administering staphylococci into the peritoneal cavity has led to the widespread use of this model and the identification of capsular polysaccharides as a key virulence determinant ( 147 ). Several studies indicate however that α-toxin is the key determinant for the lethal outcome of intraperitoneal S. aureus challenge ( 43 , 148 ). In agreement with this, active immunization with Hla H35L or passive immunization with Hla-neutralizing antibody protects mice against the lethal outcome of intraperitoneal S. aureus inoculation but does not affect intraperitoneal abscess formation ( 43 , 149 ).
S. aureus infection of the lower respiratory tract leads to significant morbidity and mortality ( 150 ). Disease severity correlates with staphylococcal isolates in particular CA-MRSA strains ( 151 ). S. aureus is also a frequent cause of ventilator-assisted pneumonia in patients with significant co-morbidities ( 152 ). Instillation of staphylococcal suspensions (4×10 8 CFU S. aureus Newman) into the left nare of anesthetized adult mice that are held upright causes labored breathing marked by a high respiratory rate and exaggerated chest wall excursion immediately after infection ( 45 ). This state resolves within 6 h, however the combined effects of aspiration and S. aureus virulence leads to the development of pneumonia with 50% mortality at 24 h, followed by an additional 20% of the animals succumbing within 48 h following inoculation ( 45 ). At these time points, all infected animals appear ill, and have an increased respiratory rate, hunched posture, and decreased mobility. A small reduction of the inoculum (8×10 7 CFU) results in no mortality, although all animals appear ill but recover fully within 48 h. Inoculation with 8×10 8 CFU of S. aureus Newman results in about 90% mortality by 24 h. For bacterial enumeration (CFU) and histopathology analysis of lungs, animals may be infected with 2×10 8 CFU of S. aureus ( 45 ). Histopathology analysis reveals infiltration of immune cells and destruction of alveolar architecture by 24 h post challenge. These damages correlated with increased replication of staphylococci ( 45 ). Animals that survive beyond the 72 h observation period, resolve the infection, show lungs with restored air space and only residual inflammation on alveolar walls ( 45 ). α-hemolysin is the key virulence factor for the pathogenesis of S. aureus pneumonia in this model, whereas surface proteins and other secreted product play either no specific or minor roles ( 45 , 60 ). Mutant mice that lack expression of the ADAM10 receptor for α-hemolysin display resistance towards S. aureus pneumonia ( 63 ). Both active (Hla H35L ) and passive immunization of mice with neutralizing antibodies provides complete protection against lethal pneumonia following challenge with both MSSA and MRSA strains ( 60 , 153 ). In a pneumonia model with neonatal mice, SpA has been shown to function as a virulence factor ( 154 ). In this model, SpA activates tumor necrosis factor receptor 1 (TNFR1) and epidermal growth factor receptor (EGFR) signaling cascades, thereby perturbing tight junctions of alveolar epithelia ( 155 , 156 ). Compared to wild-type S. aureus, a spa mutant fails to transmigrate the epithelial monolayer in tissue culture media ( 157 ). tnfr-1 null mice were found to be more resistant to pulmonary infection as compared to wild-type animals, providing support for a distinct host pathogen interaction mediated by SpA ( 158 ).
Models requiring surgery
S. aureus frequently infects medical implants and catheters by forming biofilms on the surfaces of these inert structures ( 1 , 159 ). Several different mouse models involving S. aureus infection of surgically implanted medical devices and catheters have been developed ( 160 - 163 ). Because of space constraints, we must limit the discussion below to two mouse models involving surgery: staphylococcal osteomyelitis and endocarditis.
Osteomyelitis or musculoskeletal infection of children occurs with an annual incidence of 1 in 5,000-10,000 in the United States ( 164 , 165 ). In adults, osteomyelitis is associated with compound fractures, contiguous soft-tissue infections, and diabetes ( 166 ). As with other staphylococcal infections, CA-MRSA is a causative agent of osteomyelitis in otherwise healthy individuals. S. aureus inflicts significant bone destruction and local inflammation, and, if not controlled, disseminates into the blood stream. Thus, treatment of osteomyelitis involves surgical debridement followed by prolonged antimicrobial therapy and is complicated by bacteremia, venous thrombosis, and pathologic fractures ( 167 ). A mouse model of osteomyelitis was recently described whereby a 2-μl suspension containing 10 6 CFU of S. aureus is inoculated into the intramedullary canal of the femurs of animals following the creation of a 1-mm unicortical bone defect ( 168 ). For this procedure, a small incision is made to expose the left femur of the hind limb of anesthetized animals, and trephination with a 21-gauge needle is performed to create a 1-mm diameter unicortical bone defect at the mid-femur. Bacteria are delivered through the bone defect into the intramedullary canal. All exposed areas are immediately closed with suture, and mice administered buprenorphine every 12 h for 72 h following surgery ( 168 ). Disease has been assessed by CFU enumeration upon removal of femur. Cassat and colleagues also used high-resolution microcomputed tomography and histology for visualization of bone loss and abscess formation ( 168 ). Continuous observation over 14 days shows that the staphylococcal burden peak at day 4 and imaging analysis of microcomputed tomography reveals profound bone remodeling (both destruction and formation) at the damaged site. Infected femurs lose up to 50% of their original cortical volume. Mock-infected animals heal rapidly following surgery. Histopathology examination demonstrates substantial abscess formation throughout the bone marrow of infected femurs. A mutant lacking sae was found to be attenuated in this model ( 168 ). The sae locus encodes a two component regulatory system that controls the secretion of numerous proteins in S. aureus. The Sae-regulated protease aureolysin was found to be a key contributor for this disease by mediating the degradation of PSMs, which in turn triggered osteoblast cell death and bone destruction ( 168 ). Of note, aureolysin is not a virulence factors for S. aureus induced septic arthritis ( 169 ).
Infective endocarditis (IE) occurs in 30-60% of patients with S. aureus bacteremia and is associated with 40-50% mortality ( 170 ). IE is characterized by the formation of septic vegetations on the endocardium that consist of a meshwork of host factors, such as fibrin and platelets, as well as bacterial aggregates ( 171 ). Currently, S. aureus is the leading causative agent of infective endocarditis, a rapidly progressing, destructive infection of the heart valves ( 172 , 173 ). An increased frequency of nosocomial endocarditis caused by S. aureus has also been documented, possibly linked to increased use of intravascular devices ( 172 ). It has also been reported that S. aureus bacteremia in patients with intravascular devices often results in venous thrombosis ( 174 ). A robust animal model of S. aureus endocarditis should feature aortic valve bacterial vegetations resembling those in humans. These features have been achieved in a mouse model of experimental infective endocarditis, which encompasses intravenous challenge with 10 6 CFU S. aureus one day following aortic valve trauma ( 175 ). Valve trauma is produced by introduction of an indwelling 32-gauge polyurethane catheter (sealed) into the aortic valve via the left carotid artery on anesthetized animals ( 175 ). Animals are placed in dorsal recumbency, the hair on the ventral neck and chest region shaved, and the shaved skin cleaned. Animals are kept on a vented warming table for thermoregulation and an incision is made from the thoracic inlet to the ramus of the left mandible. The left carotid artery is isolated and ligated cranially with suture. A surgical microscope is used to produce a small hole in the carotid artery with a 27-gauge needle to insert of the sealed tube (catheter) that is fed retrograde through the aortic valve (for a length of 1.3-cm). The catheter is secured in place by sutures and the incision closed. Catheters are left in place for the duration of the study and mice are infected 18 to 24 h following this procedure ( 175 ). The model has been validated by using a vancomycin regimen at exposures comparable to human dosing ( 175 ). It has also been used to demonstrate the essential contribution of staphylocoagulases, Coa and vWbp, to IE ( 42 ). Because of the small size of the carotid artery, the mouse is a technically challenging model for the study of IE, which can also be examined in rats or rabbits ( 176 , 177 ).
Conclusions and Perspectives
Mouse models have been essential for the study of several different infectious diseases caused by S. aureus, revealing virulence determinants and supporting efficacy studies with vaccine antigens or immune therapeutics. This work demonstrated also that S. aureus infection does not elicit protective immune responses against subsequent challenge with the same strain or with different S. aureus isolates ( 112 , 178 ). These insights initiated a search for immune evasive factors that may represent key targets for vaccine development, however only some of these targets can currently be studied in mice ( 114 ). These limitations may be addressed by developing transgenic animals that express human factors critical for disease pathogenesis, as has been achieved for IsdB-mediated heme-iron scavenging in mice transgenic for human hemoglobin ( 179 , 180 ). Assessing the contributions of several other human-specific virulence factors, for example SCIN, CHIPS, and SAK, which directly or indirectly inhibit C3 convertase, C5a receptor, and C3b, will require development of new transgenic models ( 49 , 181 ). Another key insight is the realization that different types of diseases (sepsis, abscess formation, peritonitis, osteomyelitis, etc.) require different types of virulence factors. Thus, if key virulence determinants were chosen as vaccine antigens, single antigen vaccine approaches cannot be expected to address all of the staphylococcal diseases discussed herein. A universal vaccine, derived from S. aureus virulence factors, would need to incorporate several different antigens into a single vaccine formulation ( 3 , 101 , 119 , 178 , 182 ). Finally, mouse models, developed for any one S. aureus disease, can reveal virulence contributions and vaccine efficacy of a specific antigen(s), and this information may be useful to predict the success of human clinical trials for the same disease. It seems unlikely, however, that such data may predict the success of clinical trials addressing another disease. If so, studies with capsular polysaccharide vaccines in the mouse peritonitis model cannot predict vaccine success in patients with end-stage renal disease and hemodialysis-related bacteremia ( 123 , 183 ). Further, studies with IsdB in mouse models for sepsis or abscess formation are unlikely to predict IsdB vaccine success in patients with mediastinal infections following cardiothoracic surgery ( 102 , 103 ). We presume that mouse models for S. aureus diseases will remain the most important surrogates for the study of staphylococcal infections in humans, their therapy and prevention.

Mouse models for S. aureus infections. Mice are susceptible to S. aureus infections. Staphylococci can be administered by four different routes: intravenous, intraperitoneal, subcutaneous and intranasal inoculation. Intravenous delivery of staphylococci generates metastatic infectious lesions in multiple internal organs including heart and kidneys. In heart, agglutinated staphylococci (staphylococcal abscess community; SAC) are surrounded by necrotic cardiac myocytes (red). In renal tissue, SAC are surrounded by fibrin deposits (brown - eosinophilic pseudocapsule) which separate bacteria from massive immune cell infiltrates (purple). Injection of staphylococci into the peritoneum creates a lesion comprised of a large number of immune cells with staphylococci. This lesion is attached to the peritoneal lining and surrounded by an inner layer of fibrin deposits (brown) and an outer layer of collagen (blue). Subcutaneous injection of staphylococci generates a subcutaneous abscess and dermonecrotic (red) lesions on the overlying skin. Lower respiratory tract infection caused by intranasal inoculation of staphylococci is characterized by obstruction of airspace (red) by inflammatory cell infiltrates and aggregates of S. aureus.
Summary of mouse models for S. aureus infection
| Disease model | Infection route | Infectious dose (CFUs) | Phenotype | References |
|---|---|---|---|---|
| Skin infection | Subcutaneous | 1×10 – 1×10 | Dermonecrosis caused by secreted toxins | ( , ) |
| Metastatic abscess formation | Intravenous | 1×10 – 1×10 | Abscess formation in most internal organs | ( ) |
| Sepsis | Intravenous | 5×10 – 5×10 | Acute lethal disease within 48 hours of infection; formation of multiple lesions in heart | ( , ) |
| Peritonitis | Intraperitoenal | 5×10 (LD ) 6×10 (LD ) | Acute lethal disease within 12 hours of infection; formation of abscess lesions on peritoneal surfaces | ( ) |
| Pneumonia | Intranasal | 2-4×10 | Acute lethal disease within 72 hours of infection; infiltration of inflammatory cells into alveolar air space | ( ) |
- Staphylococcus aureus is a human commensal
- S. aureus causes invasive and recurrent diseases
- S. aureus infections fail to elicit protective immune responses
- S. aureus disease attributes can be studied in experimental mouse models of infection
ACKNOWLEDGEMENTS
Work on staphylococcal diseases in our laboratory was supported by grants from the National Institute of Allergy and Infectious Diseases (NIAID), Infectious Diseases Branch (AI038897, AI052474, AI075258). The authors acknowledge membership within and support from the Region V “Great Lakes” Regional Center of Excellence in Biodefense and Emerging Infectious Diseases Consortium (NIH Award 1-U54-AI-057153).
Competing interests: The authors declare a conflict of interests as inventors of patent applications that are related to the development of Staphylococcus aureus vaccines and are currently under commercial license.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
- Open access
- Published: 07 September 2024
Oral administration of Lactobacillus plantarum expressing aCD11c modulates cellular immunity alleviating inflammatory injury due to Klebsiella pneumoniae infection
- Yang Zeng 1 ,
- Tiantian Li 1 ,
- Xueyang Chen 1 ,
- Xiaowei Fang 2 ,
- Chun Fang 1 ,
- Xiongyan Liang 1 ,
- Jing Liu 1 &
- Yuying Yang 1
BMC Veterinary Research volume 20 , Article number: 399 ( 2024 ) Cite this article
Metrics details
Klebsiella pneumoniae (KP), responsible for acute lung injury (ALI) and inflammation of the gastrointestinal tract, is a zoonotic pathogen that poses a threat to livestock farming worldwide. Nevertheless, there is currently no validated vaccine to prevent KP infection. The development of mucosal vaccines against KP using Lactobacillus plantarum ( L. plantarum ) is an effective strategy.
Firstly, the L. plantarum strains NC8-pSIP409-aCD11c’ and NC8-pLc23-aCD11c were constructed via homologous recombination to express the aCD11c protein either inducibly or constitutively. Both NC8-pSIP409-aCD11c’ and NC8-pLc23-aCD11c strains could enhance the adhesion and invasion of L. plantarum on bone marrow-derived dendritic cells (BMDCs), and stimulate the activation of BMDCs compared to the control strain NC8-pSIP409 in vitro. Following oral immunization of mice with NC8-pSIP409-aCD11c’ and NC8-pLc23-aCD11c, the cellular, humoral, and mucosal immunity were significantly improved, as evidenced by the increased expression of CD4 + IL-4 + T cells in the spleen, IgG in serum, and secretory IgA (sIgA) in the intestinal lavage fluid (ILF). Furthermore, the protective effects of L. plantarum against inflammatory damage caused by KP infection were confirmed by assessing the bacterial loads in various tissues, lung wet/dry ratio (W/D), levels of inflammatory cytokines, and histological evaluation, which influenced T helper 17 (Th17) and regulatory T (Treg) cells in peripheral blood and lung.
Conclusions
Both the inducible and constitutive L. plantarum strains NC8-pSIP409-aCD11c’ and NC8-pLc23-aCD11c have been found to stimulate cellular and humoral immunity levels and alleviate the inflammatory response caused by KP infection. These findings have provided a basis for the development of a novel vaccine against KP.
Peer Review reports
Klebsiella pneumoniae (KP) is a common zoonotic pathogen located at the respiratory and gastrointestinal tracts, leading to acute lung injury (ALI) and gastrointestinal infections [ 1 , 2 , 3 , 4 ]. Since the initial isolation of which in airway secretions in 1875, KP has exhibited an increasing resistance to the external environment, which results in challenges due to its drug resistance and virulence [ 5 ]. With the increasing isolation rate of KP in the global livestock farming industry, which has raised increasing concerns regarding the food safety and economic implications [ 6 , 7 ]. Bacterial vaccines have shown efficacy in reducing pathogenic bacterial infections, while the complex structural composition and numerous serotypes of KP have hindered the development of targeted commercial vaccines [ 8 , 9 , 10 ]. As one of the most predominant conditional pathogens, KP primarily causes disease through the mucosal route of infection, particularly in instances of compromised host immunity [ 11 ]. Consequently, oral vaccinations are feasible to enhancing the mucosal immunity and preventing pathogens such as KP [ 12 ].
Lactobacillus , an edible beneficial microorganism, is involved in regulating the dynamic equilibrium of intestinal flora and promoting the proliferation of immunological cells to modulate the immune response [ 13 ]. Research on Lactobacillus -based live vector vaccines has advanced significantly across various pathogens. In recent years, different expression systems have been widely utilized in Lactobacillus spp., including the constitutive or inducible expression vectors. For example, the constitutive expression vector pOri23, which was based on the P23 promoter modification, and the vector pSIP409, which was based on the sppK, sppR inducible expression system, have facilitated precise manipulation of the expressed target genes [ 14 ]. Therefore, we replaced the inducible expression system in the pSIP409 vector with the P23 promoter to construct the constitutive expression vector pLc23. Since the crucial role of antigen-presenting cells (APCs), such as dendritic cells (DCs), in antigen presentation and uptake, the use of recombinant vectors fused with receptor molecules targeting DCs can significantly enhance their effectiveness in order to enhance the uptake of antigens by APCs during immunization [ 15 ]. In a previous study, Lactobacillus plantarum ( L. plantarum ) strain NC8-pSIP409-aCD11c was employed to expresses a single-chain antibody against CD11c (scFv-CD11c, aCD11c), which efficiently bound DCs, induced DC maturation, promoted T cell differentiation, and enhanced B cell production in vivo [ 16 ].
In the development of L. plantarum vaccines, it is imperative to thoroughly understand the immunomodulatory mechanisms of L. plantarum . In case studies related to respiratory diseases, changes in T helper 17 (Th17) and regulatory T (Treg) cell numbers are critical for disease progression and are linked to COPD, lung cancer, and tuberculosis [ 17 , 18 , 19 ]. The stability and suppression of CD4 + CD25 + Treg cells depend on FOXP3, a transcription factor, and FOXP3 expression and regulation require phosphorylated STAT5 (p-STAT5) [ 20 ]. Consequently, this pathway regulates the prevention of Treg’s immunological inflammation. It has been shown that the STAT5/FOXP3 signaling pathway was dramatically suppressed in a mouse model of asthma, increasing Th17 cells and decreasing Treg cells in the bronchoalveolar lavage fluid (BALF) [ 21 , 22 , 23 ]. Therefore, in this study, the inducible and constitutive L. plantarum expressing aCD11c (NC8-pSIP409-aCD11c’ and NC8-pLc23-aCD11c) were used to investigate the effect of aCD11c expression on DCs activation, and to elucidate the relationship between the immunomodulatory effects of L. plantarum on KP infection and the activation of the STAT5/FOXP3 signaling pathway.
Materials and methods
Strains, plasmids, and primers.
Table 1 lists the strains, plasmids, and primers used in this study. L. plantarum NC8 (CCUG 61730) [ 24 ] was kindly provided by Prof. Chunfeng Wang (Jilin Agricultural University, China). The NC8 strains were quiescently grown in anaerobic conditions at 37 °C in an MRS medium containing erythromycin (Em) (10 µg/mL). E. coli Top10 strain was cultured in Luria–Bertani (LB) broth under shaking conditions at 37 °C (200 µg /mL Em). The K. pneumoniae HRB2020005 strain isolated from swine was kindly provided by Prof. Liancheng Lei (Jilin University, China) and identified via 16s rDNA (GeneBank OQ674507).
aCD11c-expressing L. plantarum Construction
The plasmid extracted from Top10-pSIP409-aCD11c was utilized as a template. The aCD11c fragment was amplified using two primer pairs, 409-a’-F/409-a’-R and pLc23-a-F/409-a’-R (Table 1 ), respectively. Subsequently, the fragments were ligated with vectors pSIP409 ( Hind III) and pLc23 ( Hind III) using the Seamless Assembly Cloning Kit (Clone Smarter Technologies, China) to generate the recombinant plasmids pSIP409-aCD11c’ and pLc23-aCD11c. After sequencing and verification by Biocorp (Tsingke Biotechnology, China), the recombinant plasmids were electro-transformed into L. plantarum NC8 using an electroporation gene introducer (Bio-Rad, USA) with parameters set at 2.5 KV, 400 Ω, and 25 µF. This resulted in the generation of recombinant L. plantarum strains NC8-pSIP409-aCD11c’ and NC8-pLc23-aCD11c, abbreviated as 409-a and pLc23-a, respectively.
Western blotting of aCD11c
The recombinant strains were inoculated in an MRS medium for anaerobic culture. SppIP (50 ng/mL, sakacin P) was added to the NC8-pSIP409 (409) and 409-a to induce protein expression, except the strain pLc23-a. After incubation, the above three strains were harvested via centrifugation, and protein samples were obtained as previously described [ 16 ]. The samples were evaluated by western blotting using an HA-labeled primary antibody (1:1000, Beyotime, China), followed by an HRP-conjugated goat anti-mouse IgG (1:5000, Solarbio, China) as the secondary antibody. Detection was performed using a chemiluminescence imager.
Adhesive and invasive ability of recombinant L. plantarum to BMDCs
C57BL/6 mice (5–6 weeks old) were obtained from the Experimental Animal Center of Three Gorges University, Yichang. Bone marrow-derived dendritic cells (BMDCs) were acquired according to the previously methods [ 25 ]. BMDCs were isolated from the tibiae and fibulae of mice. The culture medium of BMDCs was supplemented with 20 ng /mL GM-CSF and 10 ng /mL IL-4 (PeproTech, USA). On the 8th day, cells were harvested and placed in 24-well culture dishes at a density of 2 × 10 5 cells per well. The cells were then incubated for 24 h. Following that, the adhesion and invasion test of 409-a and pLc23-a were conducted. The strains were cultivated together with cells (MOI = 1000), and the monoclonal anti-mouse CD11c antibody (Bioss, China) was introduced. Following a two-hour period of stimulation, the cells were exposed to aseptic PBS solution containing 0.2% Triton X-100 for a duration of 10 min. After being diluted in a gradient manner, then the cells were incubated in MRS culture plates (37 °C, 10 µg/mL Em) overnight. Subsequently, the cells were counted in order to evaluate the rate of L. plantarum adhesion. Concurrently, those cells were exposed to L. plantarum stimulation for 2 h were subjected to invasion assays, after being treated with gentamicin (500 µg /mL). The adhesion or invasion ratio was computed according to the previously method [ 16 ].
Activation of BMDCs cells
After BMDCs were cultured in 24-well plates, 409, 409-a, pLc23-a, and LPS were added to each group of cells. L. plantarum was added to the cultures at an MOI of 10. Subsequent experiments were performed after overnight incubation. Cells were collected and incubated with antibodies purchased from BD, such as APC-labeled anti-mouse CD11c, FITC-labeled anti-mouse CD40, PerCP-Cy5.5-labeled anti-mouse CD80, and their respective isotype control antibodies. Then, all samples were processed by flow cytometry (FCM) (BD FACSMelody, USA) for analysis. The databases were parsed by FlowJo V10.
Immunization and challenge
BALB/c mice (6–8 weeks old) were supplied by the Experimental Animal Center of Three Gorges University, Yichang, Hubei Province, China. In total, 75 same-aged mice were randomly and equally categorized into five groups, namely, PBS, KP, 409, 409-a, and pLc23-a. Mice were immunized orally twice consecutively on 1st to 3rd day and 15th to 17th day (Fig. 1 A). The L. plantarum immunization dosage was 10 9 CFU/100 µL/ mouse. Furthermore, the PBS group was fed an equal volume of PBS. One week after the immunization, 3 mice were randomly selected from the PBS and L. plantarum groups to perform FCM assays.
On the 25th day, all BALB/c mice, except those in the PBS group, were injected with 1 × 10 7 CFU KP by intraperitoneal injection. After 24 h, the lung, spleen, and liver organs were removed and minced from euthanized mice after rapid cervical dislocation. The tissue homogenates were appropriately diluted and incubated on LB agar plates in a 37 °C incubator for colony counting. Colonization was calculated as the ratio of organ colonies to inoculation colonies.

Detection of CD4 + T cells activation via FCM and antibody levels through ELISA. ( A ) Immunization and infection processes in mice and the implementation of separate assays. Flow chart ( B ) and analysis ( C ) of the expression of IFN-γ + CD4 + T cells and IL-4 + CD4 + T cells in spleen. ( D ) The levels of sIgA in ILF and IgG in the serum of mice after the second immunization. A one-way ANOVA test was performed to determine statistical significance (* P < 0.05, ** P < 0.01 and *** P < 0.001)
Characterization of CD4 + T cells in the spleen
On the 24th day, 3 mice per group were measured by FCM. The spleens were aseptically removed and ground in RPMI 1640 medium. Furthermore, the supernatant was discarded via centrifugation and lysed by erythrocyte lysis solution, washed with PBS, and permeabilized. Subsequently, the cells were counted after re-suspension in 1 mL of culture medium. Surface and intracellular staining of T cells were separately performed. For intracellular staining, RPMI1640 complete medium (10% FBS) containing PMA, Ionomycin and Brefeldin A stimulating agent (BD Leukocyte Activation Cocktail, USA) was added into a 48-well culture plate containing 2 × 10 6 cells per well and cultured in an incubator for 5 h (37 °C, 5% CO 2 ) for stimulation. After the completion of stimulation, BV510 labeled anti-mouse CD3e and BV421 labeled anti-mouse CD4 were added for surface staining, APC labeled anti-mouse IFN-γ and PE-Cy7 labeled anti-mouse IL-4 and their respective isotype control antibodies (BD Pharmingen) were used for intracellular staining after fixation and permeabilization (BD fixation/permeabilization solution kit, USA). Lastly, the single-cell suspended solution was re-suspended in PBS for FCM assays.
Blood samples were obtained after the second immunization, and the supernatants were collected via centrifugation. Intestinal tissues were washed with pre-cooled PBS containing 1% protease inhibitor PMSF (Beyotime, China). Furthermore, the contents were obtained, and the supernatant was collected via centrifugation. Mucosal antibody secretory IgA (sIgA) in intestinal lavage fluid (ILF) was detected as per the manufacturer’s protocol (MEIMIAN, China). Concurrently, IgG levels in the serum were detected using the Cytokine ELISA Kit for Mouse IgG (MEIMIAN, China).
Flow cytometry assays for Th17 and Treg cells after KP infection
On the 32nd day, peripheral blood of 3 mice from each group was obtained, and one-half of the anticoagulated blood was supplemented with RPMI1640 complete medium (10% FBS) containing PMA, Ionomycin and Brefeldin A stimulating agent (BD Leukocyte Activation Cocktail, USA), which was cultured in an incubator for 5 h (37 °C, 5% CO 2 ) and homogenized every 1 h. The suspensions were incubated with BV421 labeled anti-mouse CD4, fixed, permeabilized, and incubated with PE-labeled anti-mouse IL-17 A and their respective isotype control antibodies. This was then rinsed with PBS and resuspended for FCM detection of Th17 cells. Concurrently, lungs were obtained under aseptic conditions to prepare single-cell suspensions. Tissue fragments were digested in RPMI1640 digestion solution for 30 min (10% FBS, 25 U/mL DNase I, 2.0 mg/mL collagenase IV, 1.0 mg/mL collagenase I) and mixed slowly. After digestion, the lung tissue was gently ground and filtered through a cell sieve (70 μm), centrifuged, and lysed twice by adding erythrocyte lysis solution. The lysis was terminated by PBS. The supernatant was discarded via centrifugation and suspended in RPMI 1640 complete medium to obtain single-cell suspensions. BV421-labeled anti-mouse CD4 and PE-labeled anti-mouse CD25 were first treated with cell suspensions to detect Treg cells in the other half of the unstimulated peripheral blood and lung tissues. After fixation and permeabilization, APC-labeled anti-mouse FOXP3 and their respective isotype control antibodies were added for incubation and used for FCM detection.
Lung wet/dry (W/D) ratio
On the 7th day after the KP infection, lungs were removed from mice sacrificed by cervical dislocation, washed with sterile PBS, and wet weights were obtained. Lungs were then dried in a thermostat at 65 °C for 24 h, removed, and weighed. The W/D ratio was calculated to estimate the effect of recombinant L. plantarum on pulmonary edema.
Histological evaluation (HE)
The lung and intestinal tissues of mice were obtained, washed with aseptic PBS, and immobilized in 4% paraformaldehyde solution. Furthermore, tissues were treated with dehydration and inserted in paraffin wax, sectioned, colored with hematoxylin and eosin (H&E), and observed using a microscope (Leica, Germany).
Analysis of STAT5, p-STAT5, and FOXP3
Lysis buffer RIPA (Beyotime, USA) consisting of proteinase and phosphatase inhibitor compounds was added to the samples. The mixture was homogenized and the supernatant was obtained via centrifugation. For the western blotting assay, primary antibodies included STAT5, p-STAT5, FOXP3, and GAPDH (ABclonal, China), whereas secondary antibodies included HRP-conjugated goat anti-rabbit IgG (1:5000, Solarbio, China). A chemiluminescence imager was used for detection. The results were analyzed using Image J.
Quantitative RT-PCR
The expression of tumor necrosis factor-α (TNF-α), interleukin-1β (IL-1β) and transforming growth factor-β (TGF-β) mRNA from lung tissue was detected using qRT-PCR. Total RNA was extracted using an RNA extraction kit (Tsingke, China), and reversal transcription was performed by cDNA Synthesis Kit (Vazyme, China). PCR was performed as per the manufacturer’s protocol in a reaction mixture containing 2×Universal SYBR green qPCR mix (ABclonal, China). Transcripts of the indicated genes were detected on a 7500 Real-Time PCR system Real-Time PCR system (Thermo Fisher, USA). Amplifications were processed with gene-targeted primers as follows: β-actin (AY618569), forward primer (Fw) 5’- AATCGTGCGTGACATCAAAG-3’ and reverse primer (Rv) 5’- AAGAAGGAAGGCTGGAAAAGAG-3’. TNF-α (NM_013693), Fw 5’- CAGAAAGCATGATCCGCGAC-3’ and Rv 5’-TCTGAGTGTGAGGGTCTGGG − 3’. TGF-β (M13177), Fw 5’-GCTGAACCAAGGAGACGGAA-3’ and Rv 5’- GTTGGTATCCAGGGCTCTCC-3’. IL-1β (NM_008361), Fw 5’- ATGAAAGACGGCACACCCAC-3’ and Rv 5’-GCTTGTGCTCTGCTTGTGAG-3’. The conserved gene β-actin was used as an internal control.
Statistical analysis
The GraphPad Prism 6.01 software was used for statistical analysis. Data are presented as the mean ± standard error of the mean (S.E.M.) and were assessed through one-way ANOVA (Dunnett’s multiple comparison test) in at least three independent experiments. P < 0.05 was considered statistically significant.
409-a and pLc23-a expressed aCD11c protein
The plasmid pSIP409-aCD11c’ added only the HA-tag as a flag after the aCD11c sequence to allow the detection of protein expression, while the cell wall fractions were collected to detect the expression of the aCD11c protein of strains 409-a and pLc23-a. The results showed that both different types of vectors successfully expressed aCD11c protein with the same size (39 kDa) (Fig. 2 C), indicating that aCD11c was expressed by both strains. Furthermore, the expression of aCD11c protein was higher in the inducible strain 409-a than in the constitutive strain pLc23-a under the same treatment conditions.

Structural diagrams of pSIP409-aCD11c’ and pLc23-aCD11c plasmids and the detection of aCD11c expression. ( A ) Plasmid profile of pSIP409-aCD11c’. pSIP409 is a shuttle vector that can be activated by the addition of SppIP, with the sequence MAGNSSNFIHKIKQIFTHR. ( B ) Plasmid profile of pLc23-aCD11c. pLc23 is a modified version of pSIP409 that employs a continuous expression system (pOri23) to substitute for the inducible expression system (sppK, sppR). ( C ) The expression of aCD11c. M: Low molecular weight protein standard; Lane 1: NC8-pSIP409-aCD11c’; Lane 2: NC8-pLc23-aCD11c; Lane 3: empty vector NC8-pSIP409
aCD11c-expressing strains improved adhesion and invasion of BMDCs and promoted the activation
The recombinant strains expressing aCD11c protein improved the adhesion and invasion efficiency of BMDCs. Furthermore, the inducible expression of strain 409-a was more effective than the constitutive expression of strain pLc23-a in adhesion ( P < 0.01) (Fig. 3 A) and invasion rate ( P < 0.05) (Fig. 3 A). The increased adhesion rates were distinctly decreased during the competitive assay, wherein the anti-CD11c antibody was used before co-incubation with BMDCs ( P < 0.001) (Fig. 3 A). This showed that the cellular adhesion was improved via the expression of aCD11c on the surface of the strains. Similar results were observed in the invasion study ( P < 0.001) (Fig. 3 A). This suggests that the expression of aCD11c significantly increased the numbers of bacteria in BMDCs, whereas the presence of anti-CD11c antibody decreased these results. A FACS assay was performed to further analyze the activation of targeting strains to BMDCs. Both targeting strains effectively promoted the expression of CD40 ( P < 0.05, P < 0.01) and CD80 ( P < 0.001, P < 0.05) in BMDCs compared with the 409 strain as control (Fig. 3 B). These results showed that aCD11c protein promoted the activation of BMDCs in vitro.

The adhesion and invasion of aCD11c-expression strains and activation for BMDCs. ( A ) Adhesion and invasion of targeting L. plantarum to BMDCs. ( B ) The expression of CD11c + CD40 + and CD11c + CD80 + for BMDCs were analyzed by FCM. LPS was used as a positive control. A one-way ANOVA test was performed to determine statistical significance (* P < 0.05, ** P < 0.01 and *** P < 0.001)
aCD11c-expressing strains promoted the activation of CD4 + T cells and the production of humoral antibodies
In the mouse model, to further determine the T cell response induced by the strains 409-a and pLc23-a, we determined the interferon-γ (IFN-γ) and interleukin-4 (IL-4)-producing T cells in the spleen (Fig. 1 B). The results indicated that the expression of CD4 + IL-4 + T cells from the 409-a group was upregulated compared with the 409 control group after immunization ( P < 0.05). However, the expression of IFN-γ was not considerably changed (Fig. 1 C), indicating that immunization can activate CD4 + T cells and stimulate the differentiation of CD4 + T cells toward Th2 subtypes. Further, the sIgA assay was performed to evaluate the ability of gastrointestinal mucosa to resist bacterial and viral adhesion. After oral immunization with L. plantarum , sIgA levels in ILF were significantly increased in both the 409-a ( P < 0.001) and pLc23-a ( P < 0.01) groups compared with the PBS group. Furthermore, the 409-a was significantly increased compared with the 409 groups ( P < 0.01) (Fig. 1 D). IgG, which is the main antibody in the serum that exerts antibacterial activity, was measured to determine the immune response of the body. The IgG expression in serum was increased in both the 409-a ( P < 0.01) and pLc23-a ( P < 0.05) groups compared with the 409 groups (Fig. 1 D), indicating that oral immunization could effectively induce the mucosal immunity and humoral immunity.
L. plantarum expressing aCD11c alleviated the colonization ability of KP in lung
To verify the ability of mice immunized with L. plantarum to defend against KP infection, the amounts of bacteria in the lung, liver, and spleen of different groups of mice were measured on the 7th day after KP infection. The amounts of bacteria in the lungs, livers, and spleens of mice immunized with strains 409-a and pLc23-a were significantly decreased compared with those in the control KP group ( P < 0.01) (Fig. 4 ), especially in lung tissue ( P < 0.001) (Fig. 4 A). This showed that mice immunized with L. plantarum could efficiently alleviate the colonization ability of KP in different tissues, in particularly L. plantarum expressing aCD11c could efficiently alleviate the colonization ability of KP in lung.
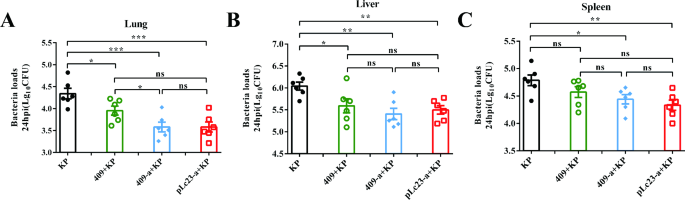
Detection of bacterial loads of different organs after KP infection for 24 h in mice., Bacterial loads of lung ( A ), liver ( B ), and spleen ( C ) organs. A one-way ANOVA test was performed to determine statistical significance (* P < 0.05, ** P < 0.01 and *** P < 0.001)
aCD11c-expressing strains reduced Th17 cells and improved Treg cells expression levels
To better evaluate the changes in the Th17 cells and Treg cells following KP infection in the mouse model, the levels of Th17 in peripheral blood and Treg cells in lung were detected 3 days after KP infection. The levels of CD4 + IL-17 A + Th17 cells in peripheral blood were significantly lower in the aCD11c-expressing group compared with those in the 409 groups ( P < 0.01) (Fig. 5 A). The 409-a + KP group and pLc23-a + KP exhibited substantially higher levels of CD4 + CD25 + FOXP3 + Treg cells than the 409 + KP group in lung ( P < 0.01 and P < 0.05) (Fig. 5 B). Furthermore, there was no significant difference in Th17 cells and Treg cells levels between the two groups 409-a and pLc23-a (Fig. 5 A and B), indicating that the expression of aCD11c reduced the Th17 cells in peripheral blood and improved Treg cells expression levels in lung.

Detection of the expression of CD4 + IL-17 A + Th17 cells and CD4 + CD25 + FOXP3 + Treg cells after KP infection. ( A ) Flow chart and analysis of the expression of CD4 + IL-17 + Th17 cells in peripheral blood. ( B ) Flow chart and analysis of the expression of CD4 + CD25 + FOXP3 + Treg cells in lung by FCM. A one-way ANOVA was performed to determine statistical significance (* P < 0.05, ** P < 0.01 and *** P < 0.001)
L. plantarum expressing aCD11c ameliorated pulmonary edema and histopathological symptoms
Following intraperitoneal injection of KP, mice in the experimental groups, as compared to the PBS group, exhibited symptoms such as a disheveled coat, restlessness, decreased appetite, reduced motility, and increased fecal excretion within 24 h. On the 7th day post-KP infection, three mice from each group were sampled, and their pulmonary tissue wet/dry (W/D) ratios were recorded. The results indicated that the W/D ratios decreased in the L. plantarum -immunized groups compared to the KP group ( P < 0.01) (Fig. 6 A), in particularly L. plantarum expressing aCD11c groups. This suggests that L. plantarum expressing aCD11c can alleviate pulmonary edema induced by KP infection. The tissues showed varying degrees of inflammatory pathological alterations, with altered lung tissue structural deformation, and inflammatory cells and exudates flooding the alveolar space. In the intestine, the intestinal villi were separated, the epithelial cells were morphologically aberrant, the intestinal mucosa was injured, and the tissue mucosa or muscle layer was destroyed. Mice in the immunized L. plantarum groups exhibited significant relief of pathological symptoms (Fig. 6 B).
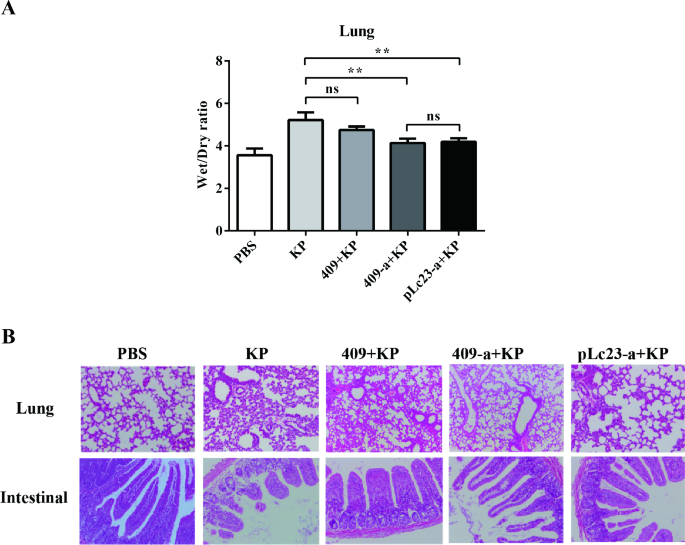
Evaluation of inflammatory response after KP infection. ( A ) W/D ratios of pulmonary tissue in each group were recorded. A one-way ANOVA test was applied to estimate statistical significance (* P < 0.05, ** P < 0.01 and *** P < 0.001). ( B ) Lung and intestinal samples from different KP-infected groups were acquired, fixed, and embedded in paraffin. Sections were stained through H&E (magnification, ×200). Scale bar, 200 μm
The targeting L. plantarum regulated Treg cells through STAT5/FOXP3 signaling pathway
KP infection can affect the inflammatory process in the lungs via NF-κB and other validation-related signaling pathways. The question arises whether targeting L. plantarum can influence the expression of Treg cells through the modulation of the STAT5/FOXP3 signaling pathway, as evidenced by the subsequent findings. The results showed an upregulation of p-STAT5 levels in the lung tissues of the 409-a and pLc23-a groups compared to the 409 group. The upregulation was notably more significant in the 409-a group after KP infection ( P < 0.001) (Fig. 7 B). Furthermore, there was significantly higher in FOXP3 expression in the 409-a ( P < 0.01) and pLc23-a ( P < 0.01) groups when compared with the 409 control group (Fig. 7 C). Analyzing the FCM results, it was observed that the levels of CD4 + CD25 + FOXP3 + Treg cells were significantly higher in lung tissue in the aCD11c-expressing groups compared to the 409 group ( P < 0.01) (Fig. 7 B). This indicates that aCD11c-expressing strains have the potential to improve Treg cell expression by regulating the STAT5/FOXP3 signaling pathway in the lung.

Activation of the STAT5/FOXP3 signaling pathway and expression of inflammatory factors in the lung after KP infection. ( A ) Expression of STAT5, p-STAT5, FOXP3, and GAPDH in the lungs of different groups of mice immunized with L. plantarum after KP infection. ( B and C ) The analysis of STAT5 phosphorylation level and evaluation of FOXP3 expression. ( D-F ) Gene expression of IL-1β, TNF-α and TGF-β in the lung. The housekeeping gene β-actin was selected as an internal control for normalization. A one-way ANOVA test was performed to determine the statistical significance (* P < 0.05, ** P < 0.01 and *** P < 0.001)
The expression of inflammatory cytokine was downregulated caused by the targeting L. plantarum
KP can cause pathogenic changes in lung tissue by activating several inflammatory signaling pathways and impairing the production of cytokines involved in inflammation. Upon assessing the expression levels of IL-1β, TGF-β, and TNF-α in lung tissue, the findings revealed that both the 409-a and pLc23-a groups significantly reduced the expression of KP-induced inflammatory cytokines IL-1β ( P < 0.001) (Fig. 7 D), TGF-β ( P < 0.01) (Fig. 7 E) and TNF-α ( P < 0.001) (Fig. 7 F) compared with that in the 409 group. This indicates that vaccination with L. plantarum expressing aCD11c can effectively prevent the production of inflammatory cytokines due to KP.
Lactic acid bacteria (LAB), as a representative intestinal probiotic, exhibit the ability for stable colonization in the gastrointestinal tract. LAB can stimulate the mucosal immune response by maintaining the equilibrium of intestinal microorganisms. Moreover, LAB functions as an effective oral vaccine delivery vehicle, offering immunological defense against pathogenic bacterial infections [ 13 ]. There is an important connection between the gut microbiome and the lung, known as the “gut-lung axis” (GLA). Disruptions to intestinal and pulmonary homeostasis can lead to allergic or inflammatory reactions. However, probiotics such as LAB can help regulate the microbiota and alleviate these conditions [ 26 , 27 , 28 ]. Despite the sophisticated nature of the pathogenesis and influencing factors involved in gastrointestinal and respiratory diseases, probiotics can effectively alleviate allergic or inflammatory reactions. This is accomplished by regulating microbiota when there is a disruption in intestinal and pulmonary homeostasis [ 29 ]. Probiotics can stimulate the body’s immune response, strengthen the protective function of the mucosal barrier, inhibit the invasion of pathogenic bacteria, and decrease the morbidity of respiratory or gastrointestinal diseases to some extent [ 30 , 31 , 32 ]. Specifically, the intestinal microbiota can mediate distal immune regulation in the lungs through the gut-lung axis [ 33 ]. In this study, oral administration of L. plantarum expressing aCD11c was found to have an immunoprotective effect against KP lung infection in mice. Furthermore, the investigation into the state of cellular and humoral immunity during immune protection and the regulatory mechanisms laid the foundation for the future development of KP vaccines.
Although KP infections are currently not major issues in livestock, the increasing detection of drug-resistant KP strains is concerning, especially with the growing integration of pets into human life [ 34 , 35 ]. Existing KP vaccines have limited efficacy due to the complex and variable serotypes of the pathogen [ 3 , 36 , 37 ]. Currently, the efficacy of pertinent vaccines cannot be assured, and they struggle to address the complex and variable serotypes of KP [ 8 , 38 ]. L. plantarum is a promising probiotic-based vaccine candidate. It can efficiently suppress harmful bacteria through metabolite production [ 39 , 40 ] and can effectively colonize both the gut and respiratory tract [ 41 ]. In the mouse model, L. plantarum immunization decreased organ bacterial loads, demonstrating its potential as a KP vaccine (Fig. 1 ).
Under normal conditions, the intestinal and pulmonary microbiomes maintain a dynamic equilibrium. The intestinal microbiota can control the threshold of immune activation and influence the systemic immune response. The mucosal immune system, which includes sIgA, acts as a key defense against inhaled pathogens in the respiratory and gastrointestinal tracts [ 42 ]. In this study, KP infection damaged the intestinal and lung mucosal barriers, while oral administration of L. plantarum was able to increase sIgA levels in ILF (Fig. 4 D). Ultimately, the mucosal integrity of the intestinal and lung tissues was superior to the control group, as observed in histopathological sections after KP infection in mice (Fig. 6 B).
DCs play a crucial role in antigen processing and T cell activation [ 15 , 43 , 44 ], when activated by foreign pathogens, they travel to nearby lymph nodes to transmit antigens to T cells, mobilizing them to promote the acquired immune response [ 45 ]. Recently, therapies targeting APCs have shown promise. The DC-SIGN, FcR, and CD11c receptors enable DCs to process foreign antigens and identify pathogens involved in innate immunity. As part of the host immune response, immature DCs acquire and internalize specific antigens, express costimulatory molecules, mature, and transport processed antigens to drive T-cell differentiation and B-cell generation [ 46 , 47 ]. In this study, L. plantarum strains expressing the aCD11c protein were able to more effectively target and activate DCs in vitro (Fig. 4 ). Analysis of the T-cell response showed that immunization with the L. plantarum strains NC8-pSIP409-aCD11c’and NC8-pLc23-aCD11c triggered a Th2-skewed immune response, with increased IL-4 production and B-cell antibody generation, rather than a Th1 IFN-γ response (Fig. 1 B and C). This Th2 response is more effective against extracellular pathogens such as bacteria, and is mediated by IL-4, which drives the maturation of B cells into plasma cells and increases antibody production [ 48 ]. This finding is consistent with the increased IgG level in humoral immunity (Fig. 4 D).
When immunity is compromised, the lung barrier cannot withstand external bacterial infestation or infection, and the absence of effective antibiotics against KP can complicate treatments. KP induces severe acute lung inflammation, such as ALI, causing respiratory failure or mortality [ 4 , 49 ]. The primary pathogenic mechanisms of ALI are triggered by inflammatory responses, oxidative stress, and apoptosis [ 50 ]. Th17/Treg is closely linked with the immunopathogenesis of prevalent clinical lung diseases such as tuberculosis and asthma [ 51 , 52 , 53 , 54 ]. Tregs can suppress the non-specific immunological effects of immune effector cells via direct contact, killing immune effector cells, or indirectly triggering apoptosis [ 55 , 56 ]. Furthermore, Tregs have the ability to inhibit the synthesis of inflammatory molecules by expressing high levels of galectin-1, eliminating Pseudomonas aeruginosa and KP [ 57 ]. The study found that immunization with aCD11c-expression L. plantarum led to a decrease in CD4 + IL-17 + Th17 cells in the peripheral blood (Fig. 5 ).
STAT5/FOXP3 signaling pathway is crucial for Treg cells evolution and function [ 20 , 57 , 58 ]. The study revealed that the group treated with aCD11c-expressing L. plantarum had significantly greater levels of phosphorylated STAT5 (p-STAT5) and the transcription factor FOXP3 than did the KP infection group. This finding suggested that L. plantarum intervention activated the STAT5/FOXP3 pathway under inflammatory conditions, leading to increased FOXP3 expression and enhanced Treg function (Fig. 7 A and B). However, the expression levels of p-STAT5 and FOXP3 were not positively correlated in the KP infection group compared to the control group. This may be because the KP infection affected the production or expression of other signaling pathways and cytokines, such as the PI3K/Akt/mTOR pathway and TGF-β, TLR, and hemoglobin, which can also contribute to FOXP3 expression through compensatory mechanisms [ 59 , 60 ]. Bacterial translocation and intrapulmonary immune response mobilization induce inflammatory factors such as IL-1β and TNF-α to damage lung tissue and trigger oxidative stress causing subsequent infection [ 61 , 62 ]. However, IL-1β, TNF-α, and TGF-β expression were lower in the aCD11c groups than that in the KP group. Furthermore, inflammatory pathological changes in the lung and gut decreased, indicating that L. plantarum intervention could reduce KP-induced inflammation (Fig. 7 C). Although TNF-α production in lung tissues was higher in the 409 + KP group than in the KP group, it probably due to the fact that L. plantarum 409 induced the production of some inflammatory factors as an exogenous stimulus or immunogen [ 63 ], the side effect was alleviated by L. plantarum 409-a and pLc23-a, which express the aCD11c protein, by stimulating the DCs to better regulate cellular immunity and humoral immunity.
In conclusion, this study revealed that the targeting of the L. plantarum strains NC8-409-aCD11c ’ and NC8-pLc23-aCD11c, which exhibit induced expression of aCD11c in the former and constitutive expression in the latter, could effectively improve the adhesion, invasion, and activation of BMDCs in vitro. L. plantarum strains expressing aCD11c were able to enhance cellular, humoral, and mucosal immunity in mice after oral immunization. Furthermore, L. plantarum -induced immunity was able to reduce inflammatory pathological changes in tissues by activating the STAT5/FOXP3 signaling pathway. This led to an increase in CD4 + CD25 + FOXP3 + Treg cells from the lungs and a reduction in Th17 cells from the peripheral blood after infection. No significant differences were found between the two L. plantarum strains in the experiments, except for protein expression. The study then determined the relationship between targeting L. plantarum , dendritic cells, and KP based on the improved protein expression of the strain with constitutive aCD11c expression.
Data availability
Data is provided within the manuscript.
Argyres MI. Hospital-acquired infections due to Gram-negative Bacteria. N Engl J Med. 2010;363(15):1483. https://doi.org/10.1056/NEJMc1006641 .
Article PubMed CAS Google Scholar
Bidewell CA, Williamson SM, Rogers J, Tang Y, Ellis RJ, Petrovska L, et al. Emergence of Klebsiella pneumoniae subspecies pneumoniae as a cause of septicaemia in pigs in England. PLoS ONE. 2018;13(2):e0191958. https://doi.org/10.1371/journal.pone.0191958 .
Article PubMed PubMed Central CAS Google Scholar
Yang F, Deng B, Liao W, Wang P, Chen P, Wei J. High rate of multiresistant Klebsiella pneumoniae from human and animal origin. Infect Drug Resist. 2019;12:2729–37. https://doi.org/10.2147/IDR.S219155 .
Zou B, Jiang W, Han H, Li J, Mao W, Tang Z, et al. Acyloxyacyl hydrolase promotes the resolution of lipopolysaccharide-induced acute lung injury. PLoS Pathog. 2017;13(6):e1006436. https://doi.org/10.1371/journal.ppat.1006436 .
Yan X, Zeng J, Li X, Zhang Z, Din AU, Zhao K, et al. High incidence and characteristic of PRRSV and resistant bacterial co-infection in pig farms. Microb Pathog. 2020;149:104536. https://doi.org/10.1016/j.micpath.2020.104536 .
Article CAS Google Scholar
Fuentes-Castillo D, Shiva C, Lincopan N, Sano E, Fontana H, Streicker DG, et al. Global high-risk clone of extended-spectrum β-lactamase (ESBL)-producing Klebsiella pneumoniae ST307 emerging in livestock in Peru. Int J Antimicrob Agents. 2021;58(3):106389. https://doi.org/10.1016/j.ijantimicag.2021.106389 .
Sirichokchatchawan W, Apiwatsiri P, Pupa P, Saenkankam I, Khine NO, Lekagul A, et al. Reducing the risk of transmission of critical Antimicrobial Resistance determinants from contaminated pork products to humans in South-East Asia. Front Microbiol. 2021;12:689015. https://doi.org/10.3389/fmicb.2021.689015 .
Article PubMed PubMed Central Google Scholar
Assoni L, Girardello R, Converso TR, Darrieux M. Current stage in the development of Klebsiella pneumoniae vaccines. Infect Dis Ther. 2021;10(4):2157–75. https://doi.org/10.1007/s40121-021-00533-4 .
Dell’Annunziata F, Dell’Aversana C, Doti N, Donadio G, Dal Piaz F, Izzo V, et al. Outer membrane vesicles derived from Klebsiella pneumoniae are a driving force for horizontal gene transfer. Int J Mol Sci. 2021;22(16):8732. https://doi.org/10.3390/ijms22168732 .
Zhang B, Hu D, Dou Y, Xiong L, Wang X, Hu J, et al. Identification and Evaluation of Recombinant Outer Membrane Proteins as vaccine candidates against Klebsiella pneumoniae . Front Immunol. 2021;12:730116. https://doi.org/10.3389/fimmu.2021.730116 .
Martin RM, Bachman MA. Colonization, infection, and the Accessory Genome of Klebsiella pneumoniae . Front Cell Infect Microbiol. 2018;8:4. https://doi.org/10.3389/fcimb.2018.00004 .
Vela Ramirez JE, Sharpe LA, Peppas NA. Current state and challenges in developing oral vaccines. Adv Drug Deliv Rev. 2017;114:116–31. https://doi.org/10.1016/j.addr.2017.04.008 .
Suez J, Zmora N, Zilberman-Schapira G, Mor U, Dori-Bachash M, Bashiardes S, et al. Post-antibiotic gut mucosal microbiome reconstitution is impaired by Probiotics and improved by autologous FMT. Cell. 2018;174(6):1406–23. https://doi.org/10.1016/j.cell.2018.08.047 .
Simon D, Chopin A. Construction of a vector plasmid family and its use for molecular cloning in Streptococcus lactis . Biochimie. 1988;70(4):559–66. https://doi.org/10.1016/0300-9084(88)90093-4 .
Balan S, Saxena M, Bhardwaj N. Dendritic cell subsets and locations. Int Rev Cell Mol Biol. 2019;348:1–68. https://doi.org/10.1016/bs.ircmb.2019.07.004 .
Liu J, Yang G, Gao X, Zhang Z, Liu Y, Yang X, et al. Immunomodulatory properties of Lactobacillus plantarum NC8 expressing an Anti-CD11c single-chain fv fragment. J Microbiol Biotechnol. 2019;29(1):160–70. https://doi.org/10.4014/jmb.1809.09017 .
Bernard-Raichon L, Colom A, Monard SC, Namouchi A, Cescato M, Garnier H, et al. A pulmonary Lactobacillus murinus strain induces Th17 and RORγt (+) Regulatory T cells and reduces lung inflammation in tuberculosis. J Immunol. 2021;207(7):1857–70. https://doi.org/10.4049/jimmunol.2001044 .
Jia L, Wu R, Han N, Fu J, Luo Z, Guo L, et al. Porphyromonas gingivalis and Lactobacillus rhamnosus GG regulate the Th17/Treg balance in colitis via TLR4 and TLR2. Clin Transl Immunol. 2020;9(11):e1213. https://doi.org/10.1002/cti2.1213 .
Nagai Y, Lam L, Greene MI, Zhang H. FOXP3 and its cofactors as targets of Immunotherapies. Engineering. 2019;5(1):115–21. https://doi.org/10.1186/s12876-015-0388-x .
Farrar MA, Harris LM. Turning transcription on or off with STAT5: when more is less. Nat Immunol. 2011;12(12):1139–40. https://doi.org/10.1038/ni.2163 .
Thomas R, Qiao S, Yang X. Th17/Treg Imbalance: implications in Lung Inflammatory diseases. Int J Mol Sci. 2023;24(5). https://doi.org/10.3390/ijms24054865 .
Garín MI, Chu CC, Golshayan D, Cernuda-Morollón E, Wait R, Lechler RI. Galectin-1: a key effector of regulation mediated by CD4 + CD25 + T cells. Blood. 2007;109(5):2058–65. https://doi.org/10.1182/blood-2006-04-016451 .
Passerini L, Allan SE, Battaglia M, Di Nunzio S, Alstad AN, Levings MK, et al. STAT5-signaling cytokines regulate the expression of FOXP3 in CD4 + CD25 + regulatory T cells and CD4 + CD25 - effector T cells. Int Immunol. 2008;20(3):421–31. https://doi.org/10.1093/intimm/dxn002 .
Sørvig E, Mathiesen G, Naterstad K, Eijsink VGH, Axelsson L. High-level, inducible gene expression in Lactobacillus sakei and Lactobacillus plantarum using versatile expression vectors. Microbiolog. 2005;151(7):2439–49. https://doi.org/10.1099/mic.0.28084-0 .
Liu J, Yang G, Huang H, Shi C, Gao X, Yang W, et al. Dendritic cells targeting Lactobacillus plantarum strain NC8 with a surface-displayed single-chain variable fragment of CD11c induce an Antigen-Specific Protective Cellular Immune Response. Infect Immun. 2020;88(2):e00759–19. https://doi.org/10.1128/IAI.00759-19 .
Budden KF, Gellatly SL, Wood DL, Cooper MA, Morrison M, Hugenholtz P, et al. Emerging pathogenic links between microbiota and the gut-lung axis. Nat Rev Microbiol. 2017;15(1):55–63. https://doi.org/10.1038/nrmicro.2016.142 .
Dang AT, Marsland BJ. Microbes, metabolites, and the gut-lung axis. Mucosal Immunol. 2019;12(4):843–50. https://doi.org/10.1038/s41385-019-0160-6 .
Willis KA, Stewart JD, Ambalavanan N. Recent advances in understanding the ecology of the lung microbiota and deciphering the gut-lung axis. Am J Physiol Lung Cell Mol Physiol. 2020;319(4):710–16. https://doi.org/10.1152/ajplung.00360.2020 .
Stavropoulou E, Bezirtzoglou E. Probiotics in Medicine: a long debate. Front Immunol. 2020;11:2192. https://doi.org/10.3389/fimmu.2020.02192 .
Mortaz E, Adcock IM, Folkerts G, Barnes PJ, Paul Vos A, Garssen J. Probiotics in the management of lung diseases. Mediators Iflamm. 2013:751068. https://doi.org/10.1155/2013/751068
Yang Y, Huang W, Fan Y, Chen GQ. Gastrointestinal microenvironment and the gut-lung Axis in the Immune responses of severe COVID-19. Front Mol Biosci. 2021;8:647508. https://doi.org/10.3389/fmolb.2021.647508 .
De Oliveira GLV, Oliveira CNS, Pinzan CF, de Salis LVV, Cardoso CRB. Microbiota Modulation of the gut-lung Axis in COVID-19. Front Immunol. 2021;12:635471. https://doi.org/10.3389/fimmu.2021.635471 .
Fanos V, Pintus MC, Pintus R, Marcialis MA. Lung microbiota in the acute respiratory disease: from coronavirus to metabolomics. J Pediatr Neonatal I. 2020;9(1):e090139. https://doi.org/10.7363/090139 .
Article Google Scholar
Jian W, Yuan-Yuan S, Shou-Yu G, Fei-Fei D, Jia-Yu Y, Xue-Hua W, et al. Serotype and virulence genes of Klebsiella pneumoniae isolated from mink and its pathogenesis in mice and mink. Sci Rep. 2017;7(1):17291. https://doi.org/10.1038/s41598-017-17681-8 .
Roberts DE, McClain HM, Hansen DS, Currin P, Howerth EW. An outbreak of Klebsiella pneumoniae infection in dogs with severe enteritis and septicemia. J Vet Diagn Invest. 2000;12(2):168–73. https://doi.org/10.1177/104063870001200215 .
Coletti M, Passamonti F, Del Rossi E, Franciosini MP, Setta B. Klebsiella pneumoniae infection in Italian rabbits. Vet Rec. 2001;149(20):626–7. https://doi.org/10.1136/vr.149.20.626 .
Wei H, Shen J, Pang X, Ding D, Zhang Y, Zhang B, et al. Fatal infection in human flora-associated piglets caused by the opportunistic pathogen Klebsiella pneumoniae from an apparently healthy human donor. J Vet Med Sci. 2008;70(7):715–7. https://doi.org/10.1292/jvms.70.715 .
Article PubMed Google Scholar
Lundberg U, Senn BM, Schüler W, Meinke A, Hanner M. Identification and characterization of antigens as vaccine candidates against Klebsiella pneumoniae . Hum Vaccin Immunother. 2013;9(3):497–505. https://doi.org/10.4161/hv.23225 .
Mohammadi F, Eshaghi M, Razavi S, Sarokhalil DD, Talebi M, Pourshafie MR. Characterization of bacteriocin production in Lactobacillus spp. isolated from mother’s milk. Microb Pathog. 2018;118:242–6. https://doi.org/10.1016/j.micpath.2018.03.020 .
Wyszyńska A, Kobierecka P, Bardowski J, Jagusztyn-Krynicka EK. Lactic acid bacteria–20 years exploring their potential as live vectors for mucosal vaccination. Appl Microbiol Biotechnol. 2015;99(7):2967–77. https://doi.org/10.1007/s00253-015-6498-0 .
Garcia-Gonzalez N, Prete R, Battista N, Corsetti A. Adhesion properties of Food-Associated Lactobacillus plantarum strains on human intestinal epithelial cells and modulation of IL-8 release. Front Microbiol. 2018;9:2392. https://doi.org/10.3389/fmicb.2018.02392 .
Li Y, Jin L, Chen T. The effects of secretory IgA in the Mucosal Immune System. Biomed Res Int. 2020;2032057. https://doi.org/10.1155/2020/2032057 .
Macri C, Pang ES, Patton T, O’Keeffe M. Dendritic cell subsets. Semin Cell Dev Biol. 2018;84:11–21. https://doi.org/10.1016/bs.ircmb.2019.07.004 .
Worbs T, Hammerschmidt SI, Förster R. Dendritic cell migration in health and disease. Nat Rev Immunol. 2017;17(1):30–48. https://doi.org/10.1038/nri.2016.116 .
Shapouri-Moghaddam A, Mohammadian S, Vazini H, Taghadosi M, Esmaeili SA, Mardani F, et al. Macrophage plasticity, polarization, and function in health and disease. J Cell Physiol. 2018;233(9):6425–40. https://doi.org/10.1002/jcp.26429 .
Hussain A, Rafeeq H, Munir N, Jabeen Z, Afsheen N, Rehman KU, et al. Dendritic cell-targeted therapies to treat Neurological disorders. Mol Neurobiol. 2022;59(1):603–19. https://doi.org/10.1007/s12035-021-02622-4 .
Mosca PJ, Lyerly HK, Clay TM, Morse MA, Lyerly HK. Dendritic cell vaccines. Front Biosci. 2007;12:4050–60. https://doi.org/10.2741/2371 .
Wang J, Jiang H, Yang R, Zhang S, Zhao W, Hu J, et al. Construction and evaluation of recombinant Lactobacillus plantarum NC8 delivering one single or two copies of G protein fused with a DC-targeting peptide (DCpep) as novel oral rabies vaccine. Vet Microbiol. 2020;251:108906. https://doi.org/10.1016/j.vetmic.2020.108906 .
Peleg AY, Hooper DC. Hospital-acquired infections due to gram-negative bacteria. N Engl J Med. 2010;362(19):1804–13. https://doi.org/10.1056/NEJMc1006641 .
Weng TI, Wu HY, Kuo CW, Liu SH. Honokiol rescues sepsis-associated acute lung injury and lethality via the inhibition of oxidative stress and inflammation. Intensive Care Med. 2011;37(3):533–41. https://doi.org/10.1007/s00134-010-2104-1 .
Zhang W, Liu X, Zhu Y, Liu X, Gu Y, Dai X, et al. Transcriptional and posttranslational regulation of Th17/Treg balance in health and disease. Eur J Immunol. 2021;51(9):2137–50. https://doi.org/10.1002/eji.202048794 .
Li XN, Pan X, Qiu D. Imbalances of Th17 and Treg cells and their respective cytokines in COPD patients by disease stage. Int J Clin Exp Med. 2014;7(12):5324–9. PMCID: PMC4307485.
PubMed PubMed Central Google Scholar
Singh D, Agusti A, Anzueto A, Barnes PJ, Bourbeau J, Celli BR, et al. Global strategy for the diagnosis, management, and Prevention of Chronic Obstructive Lung Disease: the GOLD science committee report 2019. Eur Respir J. 2019;53(5):1900164. https://doi.org/10.1183/13993003.00164-2019 .
Lourenço JD, Teodoro WR, Barbeiro DF, Velosa APP, Silva LEF, Kohler JB, et al. Th17/Treg-Related Intracellular Signaling in patients with Chronic Obstructive Pulmonary Disease: comparison between local and systemic responses. Cells. 2021;10(7):1569. https://doi.org/10.3390/cells10071569 .
Zhang Y, Guo J, Jia R, Treg. A promising immunotherapeutic target in oral diseases. Front Immunol. 2021;12:667862. https://doi.org/10.3389/fimmu.2021.667862 .
Barbi J, Pardoll D, Pan F. Treg functional stability and its responsiveness to the microenvironment. Immunol Rev. 2014;259(1):115–39. https://doi.org/10.1111/imr.12172 .
Dong Y, Yang C, Pan F. Post-translational regulations of Foxp3 in Treg Cells and their therapeutic applications. Front Immunol. 2021;12:626172. https://doi.org/10.3389/fimmu.2021.626172 .
Ono M. Control of regulatory T-cell differentiation and function by T-cell receptor signalling and Foxp3 transcription factor complexes. Immunology. 2020;160(1):24–37. https://doi.org/10.1111/imm.13178 .
Galgani M, De Rosa V, La Cava A, Matarese G. Role of metabolism in the Immunobiology of Regulatory T Cells. J Immunol. 2016;197(7):2567–75. https://doi.org/10.4049/jimmunol.1600242 .
Ogawa C, Tone Y, Tsuda M, Peter C, Waldmann H, Tone M. TGF-β-mediated Foxp3 gene expression is cooperatively regulated by Stat5, Creb, and AP-1 through CNS2. J Immunol. 2014;192(1):475–83. https://doi.org/10.4049/jimmunol.1301892 .
Aschner Y, Downey GP. Transforming growth Factor-β: Master Regulator of the respiratory system in Health and Disease. Am J Respir Cell Mol Biol. 2016;54(5):647–55. https://doi.org/10.1165/rcmb.2015-0391TR .
Del Valle DM, Kim-Schulze S, Huang HH, Beckmann ND, Nirenberg S, Wang B, et al. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat Med. 2020;26(10):1636–43. https://doi.org/10.1038/s41591-020-1051-9 .
Suez J, Zmora N, Segal E, Elinav E. The pros, cons, and many unknowns of probiotics. Nat Med. 2019;25(5):716–29. https://doi.org/10.1038/s41591-019-0439-x .
Download references
Acknowledgements
The authors would like to thank Prof. Chunfeng Wang in Jilin Agricultural University from China, for providing Lactobacillus plantarum NC8 and Prof. Liancheng Lei in Jilin University from China, for providing Klebsiella pneumoniae HRB2020005 strain.
This work was supported by Natural Science Foundation of Hubei Province (2021CFB173).
Author information
Authors and affiliations.
College of Animal Science and Technology, Yangtze University, Jingzhou, 434025, China
Yang Zeng, Tiantian Li, Xueyang Chen, Chun Fang, Xiongyan Liang, Jing Liu & Yuying Yang
College of Agriculture, Yangtze University, Jingzhou, 434025, China
Xiaowei Fang
You can also search for this author in PubMed Google Scholar
Contributions
YZ and JL conceived and designed research. YZ, TTL, XYC, XWF conducted experiments. YZ and JL analyzed data. CF, XYL, YYY contributed to the text editing. YZ wrote the manuscript and JL reviewed the manuscript. All authors all read and approved the manuscript.

Corresponding authors
Correspondence to Jing Liu or Yuying Yang .
Ethics declarations
Ethics approval and consent to participate.
We obtained informed consent from the Experimental Animal Center of Three Gorges University to use the animals in this study. In the experiments, we collected blood, spleen, intestinal lavage fluid, internal organs, liver and lung. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed (People’s Republic of China Ministry of Health, document NO.55, 2001), and all procedures were approved by the Ethics Committee of Health Science Center, Yangtze University (202401002). This study was conducted in accordance with the ARRIVE guidelines.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1
Rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/ .
Reprints and permissions
About this article
Cite this article.
Zeng, Y., Li, T., Chen, X. et al. Oral administration of Lactobacillus plantarum expressing aCD11c modulates cellular immunity alleviating inflammatory injury due to Klebsiella pneumoniae infection. BMC Vet Res 20 , 399 (2024). https://doi.org/10.1186/s12917-024-04248-9
Download citation
Received : 06 April 2024
Accepted : 26 August 2024
Published : 07 September 2024
DOI : https://doi.org/10.1186/s12917-024-04248-9
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- L. plantarum
- Klebsiella pneumoniae
- Mucosal vaccine
- Oral immunization
- STAT5/FOXP3
BMC Veterinary Research
ISSN: 1746-6148
- General enquiries: [email protected]
Volume 29, Number 4—April 2023
Research Letter
Experimental infection of north american deer mice with clade i and ii monkeypox virus isolates.
Cite This Article
The global spread of monkeypox virus has raised concerns over the establishment of novel enzootic reservoirs in expanded geographic regions. We demonstrate that although deer mice are permissive to experimental infection with clade I and II monkeypox viruses, the infection is short-lived and has limited capability for active transmission.
Monkeypox virus (MPXV; genus Orthopoxvirus , Poxviridae ), which causes mpox disease, is a zoonotic pathogen that is endemic in Central Africa (clade I) and Western Africa (clade II) ( 1 ). In mid-May 2022, the World Health Organization first reported an increasing number of mpox cases in nonendemic countries, most of which had no established travel links to endemic regions ( 2 ). By October 2022, the outbreak encompassed >100 countries with reported confirmed mpox cases ( 3 ).
The global spread of MPXV outside of regions in which this virus was known to be endemic raises concerns over reverse zoonotic events resulting in the establishment of novel wildlife reservoirs. Small mammals, including rodents, have previously been implicated as enzootic reservoirs of MPXV. In North America, studies have shown that prairie dogs are susceptible to MPXV infection and may serve as a potential reservoir, but data on other wild rodents are limited ( 4 ). Peromyscus species rodents have an extensive and geographically diverse host range spanning most regions across North America and are well-established reservoirs for several zoonotic pathogens ( 5 ).
We evaluated the competency of deer mice ( Peromyscus maniculatus rufinus ) as a potential zoonotic reservoir for MPXV by using representative isolates from both clades. We infected groups of 12 adult (>6 weeks of age) deer mice with 1 of 3 MPXV isolates through intranasal instillation. The isolates included a clade II human isolate from the 2022 outbreak (MPXV/SP2833) (challenge dose 10 6 PFU); a second clade II virus isolated directly from a North American prairie dog (USA-2003) (challenge dose 10 6 PFU); and a historical clade I isolate (MPXV/V79-1-005) (challenge dose 10 4 PFU). For each virus preparation, we administered the maximum challenge dose based on titration on Vero cells. On days 4 and 10 postinfection, we euthanized 3 male and 3 female mice and collected selected solid organs for analysis of viral titers using molecular assays targeting of envelope protein gene (B6R) ( 6 ) and infectious viral quantification assays. In addition, we collected oral and rectal swab specimens and tested them similarly to assess the potential for shedding.
We conducted animal studies in accordance with the Canadian Council of Animal Care guidelines and following an animal use document approved by an institutional Animal Care and Use Committee, in a Biosafety Level 4 laboratory of the Public Health Agency of Canada. We conducted fully validated molecular assays in accordance with Public Health Agency of Canada special pathogens diagnostic procedures.
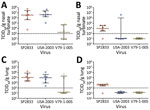

Figure . Monkeypox virus infectious titers from lung and nasal turbinate samples from experimentally infected deer mice. Groups of 12 deer mice (6 male, 6 female) were experimentally infected with monkeypox virus...
Throughout the course of the study, we observed no obvious signs of disease in any of the infected deer mice. We did not record daily weights because of the requirement for anesthetizing animals before any hands-on manipulation. Analysis of tissue samples from mice infected with the 2022 Canada isolate (MPXV/SP2833) revealed limited and sporadic spread of MPXV beyond the sites of inoculation (nasal turbinates and lungs) ( Table ). By comparison, USA-2003 appeared to disseminate beyond the respiratory tract, resulting in uniform detection of MPXV DNA in liver and spleen specimens collected at 4 days postinfection (dpi). The clade I virus (MPXV/V79-1-005) yielded results more similar to those for USA-2003; nasal turbinate, lung, liver and spleen samples were positive at 4 dpi. By day 10 dpi, organ specimens from most mice across the 3 infection groups were trending toward clearance ( Table ). Infectious titers conducted on lung and nasal turbinate specimens collected at both timepoints from the 3 challenge groups corroborated these findings and demonstrated decreasing viral titers between the 2 timepoints ( Figure ).
Of note, the clade I virus did not achieve high titers in either organ, even when analyzed at 4 dpi. Although this finding may suggest the MPXV/V79-1-005 isolate does not replicate as efficiently in deer mice, the apparent low viral titers observed may be attributable to the lower inoculum dose. A similar challenge dose of this strain resulted in lethal infection in CAST/EiJ mice ( 7 ). Further, subsequent cell culture propagations of MPXV/V79-1-005 resulted in similar titers as the clade II isolates used previously, suggesting that all 3 replicate to a similar extent on Vero cells. Nevertheless, follow-up studies with other clade I viruses are warranted.
We collected oral and rectal swab specimens to assess shedding and the potential for transmission of MPXV from infected deer mice. Overall, shedding, as suggested by the presence of MPXV DNA in swab extracts, was readily detectable in deer mice inoculated with either clade II virus at day 4, but we noted decreasing levels of positivity by day 10. Shedding of MPXV/V79-1-005 (clade 1) was far less than that of either of the clade II viruses we evaluated ( Table ).
Our study suggests that these rodents may support a short-term but abortive infection with at least clade II MPXV isolates, although with limited capacity to spread. Given the short duration of infection, these animals probably do not represent a viable enzootic reservoir for MPXV. Further studies should be conducted on other rodents in North America and Europe to assess their competency as vectors or reservoirs of MPXV. Particular interest should be given to Rattus species rodents that may frequently come into contact with medical waste containing viable MPXV.
Mr. Deschambault is a senior laboratory technician in the Special Pathogens Program of the Public Health Agency of Canada. His research interests include disease modeling and vaccine development for emerging and high - consequence viral pathogens.
Acknowledgments
This work was funded by the Public Health Agency of Canada. We obtained the MPXV USA-2003 reagent NR-2500 through the BEI Resources Repository, National Institute of Allergy and Infectious Diseases, National Institutes of Health.
All authors declare no conflict of interest.
- Likos AM , Sammons SA , Olson VA , Frace AM , Li Y , Olsen-Rasmussen M , et al. A tale of two clades: monkeypox viruses. J Gen Virol . 2005 ; 86 : 2661 – 72 . DOI PubMed Google Scholar
- Velavan TP , Meyer CG . Monkeypox 2022 outbreak: An update. Trop Med Int Health . 2022 ; 27 : 604 – 5 . DOI PubMed Google Scholar
- Centers for Disease Control and Prevention . 2022 monkeypox outbreak global map [ cited 2022 Oct 18 ]. https://www.cdc.gov/poxvirus/monkeypox/response/2022/world-map.html
- Parker S , Buller RM . A review of experimental and natural infections of animals with monkeypox virus between 1958 and 2012. Future Virol . 2013 ; 8 : 129 – 57 . DOI PubMed Google Scholar
- Barbour AG . Infection resistance and tolerance in Peromyscus spp., natural reservoirs of microbes that are virulent for humans. Semin Cell Dev Biol . 2017 ; 61 : 115 – 22 . DOI PubMed Google Scholar
- Li Y , Olson VA , Laue T , Laker MT , Damon IK . Detection of monkeypox virus with real-time PCR assays. J Clin Virol . 2006 ; 36 : 194 – 203 . DOI PubMed Google Scholar
- Warner BM , Klassen L , Sloan A , Deschambault Y , Soule G , Banadyga L , et al. In vitro and in vivo efficacy of tecovirimat against a recently emerged 2022 monkeypox virus isolate. Sci Transl Med . 2022 ; 14 : eade7646 . DOI PubMed Google Scholar
- Figure . Monkeypox virus infectious titers from lung and nasal turbinate samples from experimentally infected deer mice. Groups of 12 deer mice (6 male, 6 female) were experimentally infected with monkeypox...
- Table . Summary of PCR results of selected organs collected from deer mice experimentally infected with 3 different MPXV isolates
DOI: 10.3201/eid2904.221594
Original Publication Date: March 06, 2023
Table of Contents – Volume 29, Number 4—April 2023
| EID Search Options |
|---|
| – Search articles by author and/or keyword. |
| – Search articles by the topic country. |
| – Search articles by article type and issue. |
Please use the form below to submit correspondence to the authors or contact them at the following address:
David Safronetz, Special Pathogens Program, National Microbiology Laboratory Branch, Public Health Agency of Canada, 1015 Arlington St, Winnipeg, MB R3E 3R2, Canada
Comment submitted successfully, thank you for your feedback.
There was an unexpected error. Message not sent.
Article Citations
Highlight and copy the desired format.
| EID | Deschambault Y, Klassen L, Soule G, Tierney K, Azaransky K, Sloan A, et al. Experimental Infection of North American Deer Mice with Clade I and II Monkeypox Virus Isolates. Emerg Infect Dis. 2023;29(4):858-860. https://doi.org/10.3201/eid2904.221594 |
|---|---|
| AMA | Deschambault Y, Klassen L, Soule G, et al. Experimental Infection of North American Deer Mice with Clade I and II Monkeypox Virus Isolates. . 2023;29(4):858-860. doi:10.3201/eid2904.221594. |
| APA | Deschambault, Y., Klassen, L., Soule, G., Tierney, K., Azaransky, K., Sloan, A....Safronetz, D. (2023). Experimental Infection of North American Deer Mice with Clade I and II Monkeypox Virus Isolates. , (4), 858-860. https://doi.org/10.3201/eid2904.221594. |
Metric Details
Article views: 543.
Data is collected weekly and does not include downloads and attachments. View data is from .
What is the Altmetric Attention Score?
The Altmetric Attention Score for a research output provides an indicator of the amount of attention that it has received. The score is derived from an automated algorithm, and represents a weighted count of the amount of attention Altmetric picked up for a research output.
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
- My Bibliography
- Collections
- Citation manager
Save citation to file
Email citation, add to collections.
- Create a new collection
- Add to an existing collection
Add to My Bibliography
Your saved search, create a file for external citation management software, your rss feed.
- Search in PubMed
- Search in NLM Catalog
- Add to Search
Experimental infection of mice with the microfilariae of Onchocerca lienalis
- PMID: 7145471
- DOI: 10.1017/s0031182000055268
Microfilariae of Onchocerca lienalis were obtained from the umbilical skin of naturally infected cattle and were injected into mice. Maximum numbers of microfilariae were recovered from the skin and ears of mice when using the subcutaneous route of inoculation. Microfilariae were distributed throughout the pinna of the ear but were concentrated towards the tip where histological sections showed them to be in the dermis and adipose tissue. Using the number of parasites recovered from the ears as an index of the intensity of infection it was found that inbred CBA/H T6T6 mice were one of the most susceptible of 11 strains of mice examined. No difference in susceptibility was found between male and female CBA mice of the same age, but marked differences were demonstrated between male CBA mice of different ages. After infection with 5 000 microfilariae the recovery of parasites from the ears increased rapidly to a peak at day 35 when 10% of the inoculum was recovered, and thereafter declined up to day 242. Over a range of inoculation doses examined it was found that there was a direct, linear relationship between the number of microfilariae recovered from the ears and the number in the inoculated dose. CBA mice showed marked resistance to reinfection with microfilariae. Six days after challenge with a secondary infection recoveries of microfilariae from the ears were only 26% of the level in challenge controls and fell to 3% of the level of controls by day 35. It is concluded that the model of O. lienalis microfilariae in CBA mice shows considerable promise as a tool for research into immunological responses to skin-dwelling microfilariae, which are the principal cause of pathology in onchocerciasis.
PubMed Disclaimer
Similar articles
- Further studies on the resistance to Onchocerca microfilariae in CBA mice. Carlow CK, Muller R, Bianco AE. Carlow CK, et al. Trop Med Parasitol. 1986 Sep;37(3):276-81. Trop Med Parasitol. 1986. PMID: 3787123
- Immunity to onchocerca lienalis microfilariae in mice. I. Resistance induced by the homologous parasite. Townson S, Bianco AE, Doenhoff MJ, Muller R. Townson S, et al. Tropenmed Parasitol. 1984 Dec;35(4):202-8. Tropenmed Parasitol. 1984. PMID: 6335314
- Immunization of calves against the microfilariae of Onchocerca lienalis. Townson S, Bianco AE. Townson S, et al. J Helminthol. 1982 Dec;56(4):297-303. doi: 10.1017/s0022149x00034684. J Helminthol. 1982. PMID: 7166632
- Parasite-specific immune responses to Onchocerca lienalis microfilariae in normal and immunodeficient mice. Carlow CK, Dobinson AR, Bianco AE. Carlow CK, et al. Parasite Immunol. 1988 May;10(3):309-22. doi: 10.1111/j.1365-3024.1988.tb00223.x. Parasite Immunol. 1988. PMID: 3412786
- Clinical responses in human onchocerciasis: parasitological and immunological implications. Mackenzie CD, Williams JF, Guderian RH, O'Day J. Mackenzie CD, et al. Ciba Found Symp. 1987;127:46-72. doi: 10.1002/9780470513446.ch5. Ciba Found Symp. 1987. PMID: 3297560 Review.
- Survival of worm masses of Onchocerca ochengi in gerbils and hamsters: implications for the development of an in vivo macrofilaricide screening model. Ayiseh RB, Mbah GE, Manfo FPT, Kulu TK, Njotu FN, Monya E, Ndi EM, Tumanjong IM, Mainsah EN, Sakanari J, Lustigman S, Cho-Ngwa F. Ayiseh RB, et al. Parasitol Res. 2023 Jul;122(7):1581-1591. doi: 10.1007/s00436-023-07859-7. Epub 2023 May 8. Parasitol Res. 2023. PMID: 37154921
- Development and validation of an Onchocerca ochengi microfilarial hamster model for onchocerciasis drug screens. Mbah GE, Ayiseh RB, Cho-Ngwa F. Mbah GE, et al. BMC Infect Dis. 2016 Aug 11;16(1):404. doi: 10.1186/s12879-016-1753-2. BMC Infect Dis. 2016. PMID: 27515037 Free PMC article.
- A comprehensive, model-based review of vaccine and repeat infection trials for filariasis. Morris CP, Evans H, Larsen SE, Mitre E. Morris CP, et al. Clin Microbiol Rev. 2013 Jul;26(3):381-421. doi: 10.1128/CMR.00002-13. Clin Microbiol Rev. 2013. PMID: 23824365 Free PMC article. Review.
- Interleukin-12 modulates T-cell responses to microfilariae but fails to abrogate interleukin-5-dependent immunity in a mouse model of onchocerciasis. Hogarth PJ, Bianco AE. Hogarth PJ, et al. Immunology. 1999 Nov;98(3):406-12. doi: 10.1046/j.1365-2567.1999.00891.x. Immunology. 1999. PMID: 10583601 Free PMC article.
- Protective responses against skin-dwelling microfilariae of Onchocerca lienalis in severe combined immunodeficient mice. Folkard SG, Taylor MJ, Butcher GA, Bianco AE. Folkard SG, et al. Infect Immun. 1997 Jul;65(7):2846-51. doi: 10.1128/iai.65.7.2846-2851.1997. Infect Immun. 1997. PMID: 9199458 Free PMC article.
Publication types
- Search in MeSH
- Citation Manager
NCBI Literature Resources
MeSH PMC Bookshelf Disclaimer
The PubMed wordmark and PubMed logo are registered trademarks of the U.S. Department of Health and Human Services (HHS). Unauthorized use of these marks is strictly prohibited.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 04 September 2024
EXTL3 and NPC1 are mammalian host factors for Autographa californica multiple nucleopolyhedrovirus infection
- Yuege Huang 1 na1 ,
- Hong Mei 2 na1 ,
- Chunchen Deng 2 , 3 ,
- Wei Wang 2 ,
- Chao Yuan 2 , 3 ,
- Yan Nie ORCID: orcid.org/0000-0003-2521-8519 2 ,
- Jia-Da Li ORCID: orcid.org/0000-0002-4236-3518 1 , 4 &
- Jia Liu ORCID: orcid.org/0000-0001-9787-465X 2 , 3 , 5 , 6 , 7
Nature Communications volume 15 , Article number: 7711 ( 2024 ) Cite this article
8 Altmetric
Metrics details
- Functional genomics
Baculovirus is an obligate parasitic virus of the phylum Arthropoda . Baculovirus including Autographa californica multiple nucleopolyhedrovirus (AcMNPV) has been widely used in the laboratory and industrial preparation of proteins or protein complexes. Due to its large packaging capacity and non-replicative and non-integrative natures in mammals, baculovirus has been proposed as a gene therapy vector for transgene delivery. However, the mechanism of baculovirus transduction in mammalian cells has not been fully illustrated. Here, we employed a cell surface protein-focused CRISPR screen to identify host dependency factors for baculovirus transduction in mammalian cells. The screening experiment uncovered a series of baculovirus host factors in human cells, including exostosin-like glycosyltransferase 3 (EXTL3) and NPC intracellular cholesterol transporter 1 (NPC1). Further investigation illustrated that EXTL3 affected baculovirus attachment and entry by participating in heparan sulfate biosynthesis. In addition, NPC1 promoted baculovirus transduction by mediating membrane fusion and endosomal escape. Moreover, in vivo, baculovirus transduction in Npc1 −/+ mice showed that disruption of Npc1 gene significantly reduced baculovirus transduction in mouse liver. In summary, our study revealed the functions of EXTL3 and NPC1 in baculovirus attachment, entry, and endosomal escape in mammalian cells, which is useful for understanding baculovirus transduction in human cells.
Similar content being viewed by others
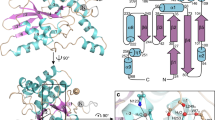
Structure of Epstein-Barr virus tegument protein complex BBRF2-BSRF1 reveals its potential role in viral envelopment
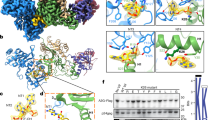
The structural basis for HIV-1 Vif antagonism of human APOBEC3G
Host neuronal PRSS3 interacts with enterovirus A71 3A protein and its role in viral replication
Introduction.
Baculoviruses are enveloped, double-strand DNA viruses comprising large genomes ranging from 80 to 180 kilobase pairs 1 . As the first baculovirus with full sequence information, AcMNPV has a genome of 133,894 nucleotides in size that contains 154 predicted non-overlapping open reading frames 2 . In nature, AcMNPV produces two structurally and functionally distinct virion forms, budded virions (BVs) and occlusion-derived virions (ODVs) 3 . BVs occur primarily in a loose viral envelope containing a single nucleocapsid, while ODVs have more than one nucleocapsid in a single envelope 4 . Although the two virion forms may share some common nucleocapsid proteins such as P39, the envelope components are largely different. BVs have envelope glycoproteins GP64, whereas ODVs contain polyhedral envelope protein P32-34, polyhedrin P29 4 , 5 , and per os infectivity factors (PIFs) that are required for the infection in insect gut 6 , 7 . BVs capture cell membranes as their envelopes and transmit within the hosts. By contrast, ODVs acquire envelopes from cell nuclear membrane and transmit between hosts 8 , 9 . BVs enter cells through receptor-mediated endocytosis 10 , 11 , while ODVs appear to fuse directly with the plasma membrane 12 , 13 .
Like other enveloped viruses, the interactions between envelope glycoproteins and host cell receptors are important for baculovirus entry. GP64 and F protein are the major envelope glycoproteins of lepidopteran nucleopolyhedrovirus (NPVs) 14 . In group I NPVs such as AcMNPV, GP64 functions as a fusion protein mediating viral entry into host cells 10 , while the F protein Ac23 seemed dispensable for infection in insect cells 14 , 15 . Different from group I NPVs, group II NPVs such as Spodoptera exigua MNPV and Lymantria dispar MNPV do not have GP64 protein and utilize F protein as the fusion protein for virus entry 16 , 17 , 18 .
As a class III fusion protein, AcMNPV GP64 protein has 512 amino acids 19 , 20 constituting five domains or structurally distinct regions, among which domain I contains fusion peptide and receptor binding peptide 10 , 19 . Different from class I and class II fusion proteins, GP64 does not undergo proteolytic cleavage 20 , 21 . The membrane fusion process of AcMNPV is triggered by a reversible, pH-dependent conformational change of GP64 22 , which remains trimeric in pre-fusion and post-fusion states 19 . Despite the existing studies in insect cells, the cellular receptor for GP64 binding in mammalian cells has not been reported.
AcMNPV is one of the most widely used baculoviruses in laboratory research and industry 23 . Baculovirus expression vector system (BEVS) was developed in the 1980s by using BVs as a transgene delivery vector to insect cells 24 . BEVS has now become one of the most widely used expression systems in the laboratory and industry for production of recombinant proteins 25 , virus-like particles 26 , 27 , and baculovirus-based vaccines 28 , 29 , 30 , 31 , 32 , 33 .
In addition to gene transfer into insect cells, baculovirus has been employed as gene delivery vectors in mammalian cells 34 , 35 , 36 . Baculovirus is capable of delivering a variety of genome editing tools into human cells, including zinc finger nucleases (ZFNs) 37 , 38 , 39 , TALENs 40 , CRISPR-Cas 41 , and CRISPR-based prime editors 42 . Genome editing in primary neurons and induced pluripotent stem cells (iPSCs) has also been achieved by baculovirus delivery of CRISPR-Cas9. Compared to other viral vectors 43 , 44 , the non-replicative and non-integrative natures render baculovirus a potentially safe gene therapy vector. Another notable advantage of baculovirus as a gene transfer vector lies in its large packaging capacity, which enables the delivery of large genome-editing tools such as prime editors in a single virion 42 .
More importantly, baculovirus has been exploited for in vivo gene delivery in mammals. Direct injection of baculovirus in mouse and rat brains could efficiently transduce neural cells 45 . Intravitreal baculovirus injection mediated gene transfer in mouse and rabbit eyes 46 , 47 . In cancer mouse models, intratumoral baculovirus injection was shown to be capable of delivering antiangiogenic proteins for cancer therapy 48 , 49 . Engineering efforts have shown that the transduction efficiency of baculovirus in mammalian cells in vitro and in vivo could be enhanced 50 , 51 , 52 , 53 .
Despite the widespread applications, the mechanism of baculovirus transduction in mammalian cells is yet poorly understood. Previous studies have shown that polybrene and heparin can inhibit baculovirus transduction in mammalian cells, suggesting an important role of electrostatic interactions 54 , 55 . Syndecan-1 (SDC-1) was also found to be important for baculovirus binding on mammalian cells 56 . Other studies suggested that baculovirus attachment on mammalian cells depended on cell surface phospholipids 57 and cholesterol 58 , 59 . Meanwhile, restriction factors that limit baculovirus transduction in mammalian cells have also been identified 60 . Nevertheless, there is yet no systematic investigation on baculovirus host factors in mammalian cells.
Genetic screen using clustered regularly interspaced short palindromic repeats (CRISPR) has been widely used in the dissection of host-pathogen interactions 61 . In a previous study, we established a cell surface protein-focused CRISPR library, so-called surfaceome CRISPR (SfCRISPR), and showed that this library could efficiently identify rhinovirus host factors 62 . In the present study, we employed this sfCRISPR library to perform forward genetic screen on a recombinant baculovirus carrying EGFP transgene (Bac-EGFP). Two rounds of negative selection identified several candidate host dependency factors for baculovirus transduction in human cells, including exostosin-like glycosyltransferase 3 (EXTL3) and Niemann-Pick C1 (NPC1). In-depth analyses illustrated the role of EXTL3 and NPC1 for the in vitro and in vivo baculovirus transduction in mammalian cells.
Identification of baculovirus host factors in HEK-293T cells using SfCRISPR screen
To identify cell surface host factors of baculovirus in mammalian cells, we performed genetic screen using a previously described cell surfaceome CRISPR library (SfCRISPR) that contained sgRNAs targeting to 1344 cell surface protein-coding genes in human genome 62 . We examined several cell lines for baculovirus transduction and found that HEK-293T cells exhibited highest transduction rate (Supplementary Fig. 1a ), which was important for reducing the background in the negative EGFP selection (Fig. 1a ). HEK-293T cells were transduced with lentivirus (LV) carrying the SfCRISPR library. Next-generation sequencing (NGS) analysis showed that this cell library had full coverage of sgRNAs (Supplementary Fig. 1b ) with a uniform distribution as predicted (Supplementary Fig. 1c ). In addition, we analyzed the shift of sgRNAs targeting essential and nonessential genes by comparing the sgRNAs at 5 days and 12 days after LV transduction and found that sgRNAs targeting essential genes were significantly depleted (Supplementary Fig. 1d ) as described 63 . These results suggested that the library was constructed successfully and was qualified for downstream phenotypic screen.
a Flowchart showing the negative selection approach for baculovirus host factor identification using surfaceome CRISPR screen. Candidate gene hits identified from the first ( b ) and second ( c ) rounds of selection. d The effects of knockout of the seven candidate genes on Bac-EGFP transduction, as determined by flow cytometry analyses. The p -value between Nontarget and EXTL3 -KO-1, EXTL3 -KO-2, NPC1 -KO-1, NPC1 -KO-2, MBTPS2 -KO-1 and MBTPS2 -KO-2 groups are 1e-5, 1e-5, 6e-6, 1e-5, 4e-5 and 4e-5, respectively. e Fluorescence imaging of Bac-EGFP infected cells. The experiment is repeated three times independently with similar results. f RT-qPCR quantification of EGFP mRNA in the lysate of EXTL3 and NPC1 knockout cells at 48 h post-Bac-EGFP transduction. β-actin is used as an internal control. g Evaluation of the effects of EXTL3 and NPC1 knockdown on Bac-EGFP transduction. The p- value between SiRNA-nontarget and SiRNA- NPC1 -2 groups is 1e-5. h Evaluation of the effects of NPC1 inhibitor U18666A on Bac-EGFP transduction. For ( d , f , and g ), Data are presented as mean ± SD ( n = 3) from three biological replicates. The significant difference is analyzed by two-tailed unpaired Student’s t- test. Mock group is the PBS treatment without baculovirus. Source data are provided as a Source Data file.
The validated cell library containing cell surface protein knockouts was then infected with baculovirus harboring enhanced green fluorescent protein transgene (Bac-EGFP). The Bac-EGFP-transduced cells were sorted by flow cytometry for enriching EGFP negative cells (Fig. 1a ). This negative selection approach allowed enrichment of cells deficient in baculovirus host factors. Two rounds of negative selection were performed and sgRNA enrichment was observed (Supplementary Fig. 1e ). With the process of negative selection, we observed an increased frequency of EGFP negative cells in round 2 selection, suggesting of successfully imposed selection pressure (Supplementary Fig. 2a, b ).
The cells after round 1 and round 2 selection were collected and the genomic DNA was extracted and analyzed by NGS. The genes with enriched sgRNAs were determined using modified robust rank aggregation (a-RRA) analyses in the MAGeCK pipeline (MAGeCK score) and candidate gene hits with an FDR cutoff of 0.05 were displayed (Fig. 1b, c and Supplementary Data File 1 ). Seven candidate hits were consistently enriched in both rounds of selection, including exostosin-like glycosyltransferase 3 (EXTL3), Niemann-Pick C1 (NPC1), SREBP cleavage activating protein (SCAP), FAM20B glycosaminoglycan xylosylkinase (FAM20B), membrane-bound transcription factor site-2 protease (MBTPS2), glucuronic acid epimerase (GLCE) and transmembrane 9 superfamily member 3 (TM9SF3). For each candidate gene, two sgRNAs were designed to construct knockout cells in HEK-293T cells (Supplementary Fig. 3a–g ).
It was found that the knockout of EXTL3 , NPC1 , SCAP , MBTPS2 and GCLE with both sgRNAs significantly reduced Bac-EGFP transduction (Fig. 1d ). Importantly, EXTL3 and NPC1 knockout reduced EGFP-positive cells to less than 50% of that in the non-targeting sgRNA group (Fig. 1d ). Thus, subsequent experiments were focused on studying the function of EXTL3 and NPC1. EXTL3 contains a glycosyltransferase domain and plays a critical role in the biosynthesis of heparan sulfate, which can facilitate viral attachment and entry 64 , 65 , 66 . NPC1 is a known host factor for various clinically important viruses 67 , 68 . Therefore, in the subsequent experiments, we focused our investigation on the functions and mechanisms of EXTL3 and NPC1 in baculovirus transduction.
Validation of EXTL3 and NPC1 as host dependency factors of baculovirus
In consistency with the flow cytometry experiments, fluorescent microscopy imaging showed that EXTL3 or NPC1 knockout reduced Bac-EGFP transduction (Fig. 1e ). Moreover, RT-qPCR analysis revealed decreased mRNA expression of EGFP transgene in EXTL3 and NPC1 knockout cells (Fig. 1f ). Similar to Bac-EGFP transduction, baculovirus carrying tdTomato (Bac-tdTomato) also exhibited reduced transduction efficiency in EXTL3 and NPC1 knockout HEK-293T cells (Supplementary Fig. 4a, b ). In addition, we designed EXTL3 - and NPC1 -targeting siRNAs (Supplementary Fig. 4c, d ) and found that knockdown of EXTL3 and NPC1 could also suppress baculovirus transduction albeit with lesser impact than EXTL3 and NPC1 knockout (Fig. 1g ). The function of NPC1 in baculovirus transduction was further validated by U18666A, a small molecule inhibitor of NPC1. It was found that U18666A inhibited Bac-EGFP transduction in a dose-dependent manner in HEK-293T cells (Fig. 1h ). Collectively, these results demonstrated that EXTL3 and NPC1 were host dependency factors for baculovirus transduction in human cells.
Evaluation of the functions of EXTL3 and NPC1 in baculovirus attachment, entry, and endosomal escape
Similar to the results in HEK-293T cells, EXTL3 and NPC1 knockout in HeLa cells (Supplementary Fig. 5a ) inhibited Bac-EGFP transduction (Fig. 2a and Supplementary Fig. 5b ). The NPC1 inhibitor U18666A reduced EGFP fluorescence in Bac-EGFP-transduced HeLa cells (Fig. 2b and Supplementary Fig. 5c ). Next we constructed single clones of HeLa cells for EXTL3 and NPC1 knockouts respectively. These single clones were confirmed to contain genomic modifications at both alleles of EXTL3 and NPC1 (Supplementary Fig. 5d ). Consistent with the above results, EXTL3 −/− and NPC1 −/− clones exhibited reduced Bac-EGFP transduction, as determined by EGFP expression at both protein (Fig. 2c and Supplementary Fig. 5e ) and mRNA (Fig. 2d ) levels. Moreover, rescue experiments by overexpressing EXTL3 and NPC1 in corresponding knockout single clones (Supplementary Fig. 5f, g ) restored baculovirus transduction (Fig. 2c, d and Supplementary Fig. 5h ).
a Evaluation of the effects of EXTL3 and NPC1 knockouts on Bac-EGFP transduction, as determined by flow cytometry analyses. The p- value between Nontarget and NPC1 -KO-2 groups is 2e-5. b Evaluation of the effects of NPC1 inhibitor U18666A on Bac-EGFP transduction. Evaluation of the effects of EXTL3 and NPC1 knockouts and overexpression rescue on Bac-EGFP transduction, as determined by flow cytometry (EGFP protein) ( c ) and RT-qPCR ( EGFP mRNA) ( d ). β-actin is used as an internal control for RT-qPCR. For ( c ), the p -values between Nontarget and EXTL3 −/− and NPC1 −/− groups are 5e-8 and 5e-6, respectively. The p -value between EXTL3 −/− and EXTL3 rescued groups is 2e-5. The p -value between NPC1 −/− and NPC1 rescued groups is 1e-5. For ( d ), the p- value between Nontarget and EXTL3 −/ − and NPC1 −/− groups are 4e-7 and 7e-8, respectively. Evaluation of the effects of EXTL3 or NPC1 knockout and overexpression rescue on Bac-EGFP attachment ( e ) and entry ( f ). g qPCR quantification of the intracellular baculovirus genome GP64 gene after Bac-EGFP transduction in non-targeting sgRNA, NPC1 −/− and NPC1 rescued HeLa cells. Bac-EGFP is incubated with cells at an MOI of 2 for 1 h and then removed from the medium. The total GP64 in each well is quantified. The p- values between NPC1 −/− and Nontarget and NPC1 rescued group at 47 h are 5e-5 and 3e-5, respectively. For ( a ), ( c – g ), Data are presented as mean ± SD ( n = 3) from three biological replicates. The significant difference is analyzed by two-tailed unpaired Student’s t- test. Mock group is the PBS treatment without baculovirus. Source data are provided as a Source Data file.
To determine the roles of EXTL3 and NPC1 during baculovirus transduction, we investigated their functions in viral attachment and entry. For viral attachment assay, Bac-EGFP was incubated with cells at 4 °C for 1 h. The cells with attached baculovirus were harvested and lysed for qPCR quantification of viral loads. For virus entry assay, Bac-EGFP was incubated with cells first at 4 °C for 1 h and then at 37 °C for 45 min to initiate the internalization of Bac-EGFP. Surface-bound Bac-EGFP was removed by trypsin treatment and the internalized virus was quantified by qPCR. It was found that EXTL3 −/− cells significantly reduced baculovirus attachment and entry, which could be restored by EXTL3 overexpression (Fig. 2e, f ). By contrast, NPC1 knockout or overexpression rescue did not significantly affect baculovirus attachment or entry (Fig. 2e, f ). Taken together, these results suggested that EXTL3, but not NPC1, was involved in baculovirus attachment and entry.
To explore whether NPC1 participated in any transduction process following attachment and entry, we sought to analyze the function of NPC1 on endosomal escape. It is known that baculovirus does not replicate in mammalian cells 69 , thus we monitored the decay rate of intracellular baculovirus genome in an approach similar to a previous study 62 . Non-targeting sgRNA-treated, NPC1 −/− and NPC1 rescued HeLa cells were transduced with Bac-EGFP for 1 h and then the virus-containing medium was replaced with fresh medium. The total GP64 quantity in each well was monitored over a course of 47 h after the removal of virus. It was found that baculovirus DNA decreased in a time-dependent manner (Fig. 2g ), suggesting that the cellular DNA degradation machinery was active toward baculovirus genome. Importantly, NPC1 −/− cells retained significantly higher level of baculovirus genome than non-targeting sgRNA-treated cells starting from 8 h after removal of virus (Fig. 2g ). Consistently, NPC1 overexpression rescue significantly reduced baculoviral genome quantity at 35 h and 47 h in comparison to NPC1 −/− cells. To exclude the possibility that the difference in intracellular viral genome was due to differential cell proliferation, we examined the proliferation of non-targeting sgRNA, NPC1 −/− and NPC1 rescued cells and observed minor difference between each group (Supplementary Fig. 5i ). These data showed that knockout of NPC1 could extend the persistence of intracellular baculovirus genome and thus indicated a potential role of NPC1 in baculovirus endosomal escape.
The above results prompted us to expand the investigation on the mechanism of retention of Bac genome. Bafilomycin A1 (BafA1) is a specific inhibitor of V-type ATPases 70 and has been reported to be capable of inhibiting endosomal escape of viruses by suppressing endosomal acidification 71 , 72 . In the present study, it was found that pre-treatment of cells with BafA1 could prolong the retention of intracellular Bac genome (Supplementary Fig. 6a ), similar to the effects of NPC1 knockout. In addition, BafA1 treatment had minor impact on cell proliferation (Supplementary Fig. 6b ), suggesting that the above-observed effects of BafA1 on intracellular Bac genome were unlikely due to its side effects on cells. Collectively, these data established a link between NPC1-mediated endosomal escape and the exposure and decay of Bac genome in cytoplasm.
EXTL3 participates in baculovirus entry through heparan sulfate biosynthesis pathway
EXTL3 (Exostosin-like protein 3) belongs to EXT protein family, which includes EXT1, EXT2, EXTL1, EXTL2, and EXTL3 73 . EXTL3 encodes a glycosyltransferase responsible for the biosynthesis of heparan sulfate (HS). Two glycosyltransferase domains, GT47 and GT64, of EXTL3 in the Golgi luminal region are the functional domains for HS biosynthesis 74 (Fig. 3a ). We constructed EXTL3 −/− HEK-293T cells (Supplementary Fig. 7a ) to explore the function of EXTL3-mediated HS biosynthesis in baculovirus transduction. It was found that baculovirus attachment and entry in EXTL3 −/− cells were markedly reduced as compared to non-targeting sgRNA cells (Fig. 3b ). These results were consistent with those in HeLa cells and illustrated an important role of EXTL3 in baculovirus attachment and entry.
a Structural organization of EXTL3 protein. b Evaluation of baculovirus attachment and entry in EXTL3 −/− cells. GP64 gene was quantified by qPCR. The p- value between Nontarget and EXTL3 −/− in entry groups is 4e-5. c Western blotting detection of heparan sulfate proteoglycan 2 (HSPG2) in wild-type and EXTL3 −/− cells. The experiment is repeated three times independently with similar results. d Analysis of the effects of heparin sodium on Bac-EGFP transduction, as determined by flow cytometry quantification of EGFP-positive cells. e Analysis of the effects of heparin sodium on Bac-EGFP attachment and entry, as determined by qPCR quantification of attached or internalized baculovirus GP64 gene. f Evaluation of the effects of HPSE on baculovirus transduction. The p -value between Nontarget and Nontarget+HPSE groups is 2e-5. g Schematic presentation of HSPG biosynthesis. h Evaluation of the effects of EXT2 knockout on baculovirus transduction. The p- value between Nontarget and EXT2 -KO groups is 7e-6. i Investigation of the relationship between EXTL3 and NPC1 functions in baculovirus transduction. The p- value between No heparin sodium and With heparin sodium in NPC1 −/− groups is 1e-6. The p- value between No heparin sodium and With heparin sodium in NPC1 rescued groups is 2e-6. For ( b , e , f , h , and i ), Data are presented as mean ± SD ( n = 3) from three biological replicates. The significant difference is analyzed by two-tailed unpaired Student’s t- test. Mock group is the PBS treatment without baculovirus. Source data are provided as a Source Data file.
Given that HS is a known baculovirus attachment and entry factor in mammalian cells 54 , 55 , 56 , we sought to explore whether EXTL3 affected baculovirus attachment and entry by affecting HS biosynthesis. We thus examined the expression of heparan sulfate proteoglycan 2 (HSPG2) protein in EXTL3 −/− HEK-293T cells and found that EXTL3 knockout abolished the production of HSPG2 (Fig. 3c ). In addition, we found that Bac-EGFP transduction could be inhibited by heparin sodium in a dose-dependent manner (Fig. 3d and Supplementary Fig. 7b ), which is a competitive inhibitor for cellular HS 65 , 75 . Similarly, 2 μg/μL heparin sodium could significantly inhibit baculovirus attachment and entry (Fig. 3e ). Moreover, 24 h pretreatment with heparanase (HPSE) could significantly reduce baculovirus transduction in non-targeting sgRNA control cells or in overexpression-rescued EXTL3 −/− cells (Fig. 3f ). These results collectively suggested that HS was important for baculovirus attachment and entry and that EXTL3 promoted baculovirus transduction by involving in HS biosynthesis.
Next, we sought to investigate whether other members in the EXT protein family have similar roles in baculovirus transduction. HS biosynthesis is initiated by the attachment of xylose to specific serine residues in HSPG core proteins, followed by the formation of a linkage tetrasaccharide, glucuronic acid-galactose-galactose-xylose (GlcA-Gal-Gal-Xyl). EXTL3 links the first N-acetyl-D-glucosamine (GlcNAc) residue to GlcA and an enzyme complex composed of EXT1 and EXT2 adds GlcA-GlcNAc disaccharide repeats to the nascent chain, followed by a series of processing reactions 76 on the chain (Fig. 3g ). We constructed EXT2 knockout HEK-293T cells (Supplementary Fig. 7c ) and found that EXT2 knockout could also affect baculovirus transduction (Fig. 3h ) though the degree of inhibition was not as prominent as those with EXTL3 knockout. These results suggested that different moieties or forms of HSPG may have differential effects on baculovirus transduction.
Next, we sought to explore the relationship between EXTL3 and NPC1 functions in baculovirus transduction. We found that in the presence of heparin sodium, the efficiency of baculovirus transduction in NPC1 −/− HeLa cells was further reduced (Fig. 3i and Supplementary Fig. 7d ). In addition, NPC1 overexpression rescue did not eliminate the inhibitory activity of heparin sodium on baculovirus transduction (Fig. 3i and Supplementary Fig. 7d ). These results suggested that NPC1 could affect baculovirus transduction through a EXTL3-independent pathway and that the functions of NPC1 and EXTL3 in baculovirus transduction were additive to each other.
NPC1 participates in baculovirus endosomal escape mainly through I and C domain-mediated membrane fusion
To elucidate the mechanism of action of NPC1 in baculovirus endosomal escape, we first used DiOC18 dye to label baculovirus particles (DiOC18-Bac-EGFP). The fluorescence of DiOC18 is self-quenched in labeled viruses and dequenched when membrane fusion occurs 77 , 78 . DiOC18-staining experiments showed that the membrane fusion of baculovirus was significantly suppressed in NPC1 −/− HeLa cells which could be restored by NPC1 overexpression, as analyzed by confocal microscopy and flow cytometry experiments (Fig. 4a, b ). These results are consistent with the above findings and strongly suggested that NPC1 promoted baculovirus endosomal escape by involving in membrane fusion.
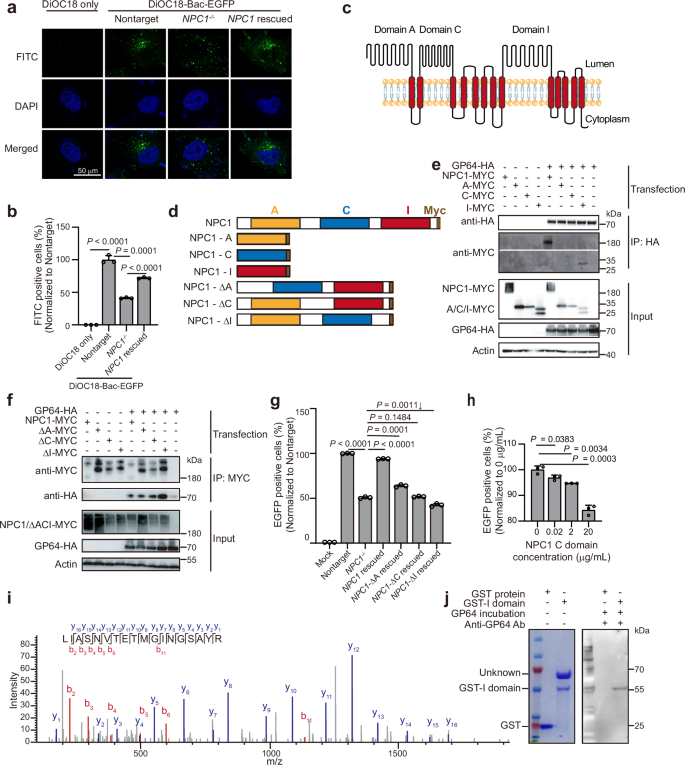
DiOC18-staining experiment for illustration of NPC1 function in baculovirus membrane fusion, as analyzed by confocal microscopy ( a ) or flow cytometry analyses ( b ). The p- value between DiOC18 only and Nontarget groups is 1e-5. The p -value between NPC1 −/− and NPC1 rescued groups is 4e-6. c The structural organization of NPC1. d Design of NPC1 constructs. Analyses of the interactions between HA-tagged GP64 and Myc-tagged full-length NPC1, NPC1-A, NPC1-C, NPC1-I ( e ), NPC1-ΔA, NPC1-ΔC or NPC1-ΔI ( f ). g Investigation of the effects of overexpression rescue with full-length or truncation constructs of NPC1 on baculovirus transduction, as determined by the EGFP-positive cells via flow cytometry. Mock, PBS treatment without baculovirus. The p -values between NPC1 −/− and Nontarget and NPC1 rescued groups are 8e-8 and 1e-7, respectively. h Neutralization of Bac-EGFP transduction in HEK-293T cells by recombinant protein of NPC1 C domain, as determined by flow cytometry analyses. i ESI-MS confirmation of purified GST-I protein ( n = 1). j Far-western analysis of the interaction between GP64 and GST-I. For ( b , g , and h ), Data are presented as mean ± SD ( n = 3) from three biological replicates. The significant difference is analyzed by two-tailed unpaired Student’s t- test. For ( e , f , and j ), The experiment is repeated three times independently with similar results. Source data are provided as a Source Data file.
NPC1 contains 13 transmembrane helices (TM) and 3 distinct lumenal domains A, C, and I 79 (Fig. 4c ). It has been reported that NPC1 affects the endosomal escape of Ebola virus through its C domain 67 , 80 . To assess which luminal domain of NPC1 interacts with GP64, we performed co-immunoprecipitation experiments. We designed constructs encoding individual domains of A, C, and I (NPC1-A, NPC1-C, and NPC1-I), and constructs with deletions of individual domain (NPC1-ΔA, NPC1-ΔC, and NPC1-ΔI) (Fig. 4d ). Co-IP analyses of the interactions between GP64 and NPC1-A, NPC1-C or NPC1-I showed that both NPC1-C and NPC1-I could interact with GP64 (Fig. 4e ). Additional Co-IP analyses showed that GP64 could interact with full-length NPC1 and truncation constructs NPC1-ΔA, NPC1-ΔC and NPC1-ΔI (Fig. 4f ), suggesting that the A, C or I domains might have redundant functions for GP64 binding. Collectively, these Co-IP experiments suggested that the C and I domains of NPC1 could both involve in the interactions with GP64.
To understand the roles of individual NPC1 domains in supporting baculovirus transduction, we performed a rescue experiment by generating stable cell lines harboring full-length NPC1 or truncation constructs on the basis of NPC1 −/− HeLa cells (Supplementary Fig. 8a ). It was found that the truncation construct NPC1-ΔA could partly restore the susceptibility of cells to baculovirus transduction whereas NPC1-ΔC or NPC1-ΔI did not significantly restore baculovirus transduction (Fig. 4g and Supplementary Fig. 8b ). These results were consistent with the Co-IP results in that C and I domains play more important roles in baculovirus transduction than A domain. In addition, we noted that while full-length NPC1 could restore baculovirus transduction to a level similar to that in non-targeting sgRNA group, none of the truncation constructs could achieve a comparable rescuing efficiency (Fig. 4g ). This suggested that the intact structural organization of NPC1 could be important for GP64 binding and baculovirus transduction. Collectively, the above results suggested that C and I domains played predominant roles in NPC1 binding with GP64 while the intact structure of NPC1 might be also important.
Next, we sought to investigate whether purified proteins of NPC1 C and I domains could interact with baculovirus or baculovirus proteins. We first performed a neutralization experiment with a commercial recombinant protein of NPC1 C domain (residues R372 to F622). It was found that NPC1 C domain could inhibit Bac-EGFP transduction in a dose-dependent manner in HEK-293T cells (Fig. 4h ). Because the function of NPC1-I domain was rarely reported in viral infection and no commercial source could be found, we expressed and purified GST tagged NPC1-I domain protein from BL21 (DE3) Escherichia coli cells. Despite of an unknown impurity band (Supplementary Fig. 8c ), the identity of the target protein band was confirmed by mass spectrometry (Fig. 4i ). We set a far WB assay to investigate the interaction between NPC-I and GP64, where GST-tagged NPC1-I domain was resolved by SDS-PAGE, transferred to a PVDF membrane and then probed with purified GP64 protein. It was found that NPC1-I domain could directly interact with GP64 in vitro (Fig. 4j ). Interestingly, we found that NPC1-C and NPC1-I shared 30% protein sequence similarity as analyzed by ClustalW (Supplementary Fig. 8d ). It was thus likely that NPC1-I resembled NPC1-C for virus interaction and endosomal escape, as reported in Ebola virus 67 , 80 , 81 .
Baculovirus transduction in mouse liver is dependent on NPC1
To understand baculovirus transduction in mice, we first examined the toxicity of intravenously administrated baculovirus. We administrated Bac-EGFP to C57BL/6J mice via tail vein and found that Bac-EGFP resulted in a transient loss of body weight in the first four days post injection (Supplementary Fig. 9a ), which was eventually recovered. Similarly, a transient change of blood routine was observed in Bac-EGFP-treated mice (Supplementary Fig. 9b ).
We then analyzed the vulnerability of different tissues to baculovirus transduction in wild-type mice and found that liver displayed highest degree of Bac-EGFP transduction (Fig. 5a, d ). It was noted that wild-type mice liver exhibited highest expression of Npc1 (Fig. 5b ). To investigate whether baculovirus transduction in mouse liver was related with Npc1 expression, we sought to construct Npc1 knockout mice. Unfortunately, Npc1 −/− C57BL/6J mice showed notably reduced birth rate and body weight (Supplementary Fig. 9c ). A previous study also showed that homozygous Npc1 knockout in mice resulted in reduced general health and pathologic changes in multiple organs 82 . By contrast, Npc1 −/+ mice did not show defects in birth rate or body weight (Supplementary Fig. 9c ), nor did liver show pathologic changes as compared to wild-type mice (Supplementary Fig. 9d ). Therefore, we used Npc1 −/+ heterozygous C57BL/6J mice for subsequent analyses. The gene disruption of Npc1 was validated in the mRNA levels (Fig. 5b ). Importantly, Bac-EGFP transduction in liver tissues of Npc1 −/+ mice was significantly reduced (Fig. 5c ) as compared to that in wild-type mice, suggesting that baculovirus transduction in mouse liver was dependent on Npc1 expression. It was also noted that in Npc1 −/+ mice, EGFP mRNA could not be detected in organs other than liver (Fig. 5d ).
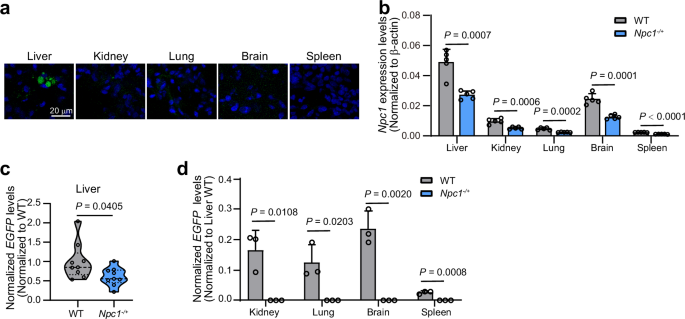
a Investigation of the tissue tropism of Bac-EGFP in wild-type mice at 72 h post intravenous injection, as determined by EGFP expression at protein ( n = 3). The fluorescence images are acquired using confocal microscopy on flash-frozen tissues that are fixed with optimal cutting temperature compound (OCT). b RT-qPCR quantification of NPC1 mRNA in the lysate of liver, kidney, lung, brain, and spleen in wild-type and Npc1 −/+ mice. The p- value between WT and Npc1 −/+ in spleen groups is 1e-8. RT-qPCR quantification of EGFP mRNA in the lysate of liver ( c ) and other tissues including kidney, lung, brain, and spleen ( d ) in wild-type and heterozygous Npc1 −/+ knockout mice at 72 h post-Bac-EGFP injection. The EGFP mRNA in the lysates of kidney, lung, brain, and spleen ( d ) in heterozygous Npc1 −/+ knockout mice is below the limit of detection. For ( b – d ), β-actin is used as an internal control. The data in this figure are from independent biological replicates. The significant difference is analyzed by two-tailed unpaired Student’s t- test. For ( b and d ), Data are presented as mean ± SD of biological replicates ( n = 5 for ( b ) and = 3 for ( d )). For c , Data are collected from nine biological replicates ( n = 9). The thick horizontal dashed line represents the median and thin horizontal dashed line indicates interquartile range between 25th and 75th percentile. Source data are provided as a Source Data file.
The large packaging capacity allows baculovirus to carry multiple transgene cassettes for the expression of large protein complexes in insect cells 83 , 84 , 85 . For the same reason, baculovirus has been an attractive gene therapy vector for delivering large or multiple gene cassettes in single viral particles, which can overcome the packaging limitation associated with conventional viral vectors. However, despite of the existing studies on baculovirus entry mechanism in mammalian cells 54 , 55 , 57 , 58 , 59 , there has been no systematic investigation on baculovirus host factors in mammalian cells. In the present study, we performed a forward genetic screen using the SfCRISPR library that carries approximately 1400 cell surface proteins. Two proteins, EXTL3 and NPC1, were identified as host dependency factors and were shown to involve in baculovirus attachment and entry, and endosomal escape processes respectively.
Investigation of EXTL3 suggested that it promoted baculovirus attachment and entry by mediating heparan sulfate (HS) biosynthesis. EXT2, another enzyme involved in HSPG biosynthesis, was also shown to affect baculovirus transduction. These results were consistent with the previous findings 54 , 55 and confirmed the importance of HS for baculovirus transduction. As HS is widely distributed across different cell types and tissues, we did not further explore the dependency of baculovirus transduction on HS. However, it is likely that different cells or tissues display different expression profiles of EXT family proteins or proteins related to HS biosynthesis, which can be important determinants for baculovirus transduction.
Moreover, it was found in the present study that NPC1 participated in baculovirus membrane fusion and endosomal escape, but not entry or attachment. These findings were consistent with the function of NPC1 in infection of filoviruses 86 and African swine fever virus (ASFV) 87 . Unlike the case in EBOV where the C domain of NPC1 interacted with EBOV glycoprotein 86 , it was found in the present study that both the C and I domains of NPC1 interacted with baculovirus GP64 protein. Additionally, an NPC1 inhibitor U18666A that can block EBOV infection was found to be efficient in inhibiting baculovirus transduction in mammalian cells. The structural basis of baculovirus GP64 interaction with NPC1-C and -I domains and the mechanism of U18666A-mediated inhibition of baculovirus transduction require further investigation. In consistency with these observations in mammalian cells, previous studies suggested that Bombyx mori NPC1 and NPC2 proteins promoted Bombyx mori nucleopolyhedrovirus (BmNPV) infection in insect cells by facilitating membrane fusion and endosomal escape 88 , 89 . More interestingly, BafA1 treatment experiments suggested the NPC1-mediated endosomal escape could release Bac genome into cytoplasm for transgene expression, the process of which will also accelerate Bac genome degradation.
One major discovery in the present study is that partial knockout of Npc1 reduced baculovirus transduction in mouse liver. To the best of our knowledge, this was the first study revealing host factors for in vivo baculovirus transduction in mammals. This result implied that the expression of NPC1 and its function in endosomal escape could be a critical determinant for baculovirus transduction in mammals. This knowledge can be exploited to design novel strategies of targeted baculovirus delivery for tissues or cells with high NPC1 expression. In addition, our study shed light on engineering baculovirus GP64 protein for enhanced interactions with mammalian NPC1. Nevertheless, although the present and previous studies revealed general safety of baculovirus administration in mice, the safety of baculovirus in humans remains unknown and should be carefully addressed in future studies.
In addition to EXTL3 and NPC1 , our studies revealed several other genes that might involve in baculovirus transduction in HEK-293T cells. Among these genes, MBTPS2 seemed to have a major impact on the transduction efficiency. MBTPS2 encodes site-2 protease (S2P), which is a hydrophobic zinc metalloprotease. S2P can cleave transmembrane proteins to release their nucleus-localizing components that regulate the transcription of genes involved in lipid biosynthesis 90 and ER stress response 91 . It has been reported that S2P can mediate the activation of antiviral proteins through regulated intramembrane proteolysis (RIP) process during HCV infection 92 . Thus, it could be interesting to investigate in future studies whether MBTPS2 plays similar functions during baculovirus transduction in mammalian cells.
One limitation in our study was that the HEK-293T cells used in the screening process were deficient of cyclic GMP-AMP synthase-stimulator of interferon genes (cGAS-STING) pathway. While the high baculovirus transduction efficiency in HEK-293T could reduce the background signal in the negative EGFP selection, the lack of cGAS-STING pathway eliminated the possibility of uncovering host factors involved in cGAS-STING signaling Recent studies showed that cGAS-STING signaling could impede baculovirus transduction in mammalian cells by affecting interferon (IFN) production 60 . Interestingly, cGAS-STING-mediated IFN production in mammalian cells could be inhibited by AcMNPV P26 protein 93 . In future studies, it would be interesting to perform CRISPR screen on cGAS-STING-competent cells to uncover baculovirus host factors that act in the context of IFN signaling.
Cell culture
HeLa cells were obtained from the American Type Culture Collection (ATCC). HEK-293T, U87-MG, Club, HK-2, U251, and A549 cells were obtained from the Cell Bank of Shanghai Institutes for Biological Science (SIBS). All the cells used in this study were validated by VivaCell Biosciences (Shanghai, China). All cells were grown in Dulbecco’s modified Eagle’s medium (DMEM, Thermo, Pittsburgh, USA) supplemented with 10% fetal bovine serum (FBS, Thermo) and 1% penicillin-streptomycin (Thermo) and maintained at 37 °C in a fully humidified incubator containing 5% CO 2. All cells were confirmed by PCR to be free of mycoplasma contamination.
Lentivirus production and transduction
To produce lentivirus (LVs), HEK-293T cells at a confluence of 70–90% were transfected with LV packaging plasmid pMD2.G, envelope plasmid psPAX, and transfer plasmid pLentiCRISPR-v2 that carried Cas9 gene and a single sgRNA or pooled sgRNA plasmid library with a mass ratio of 1: 1.5: 2 using Lipofectamine 3000 (Thermo). In the case of overexpression, the transfer plasmids were pLenti-EF1α-IRES-NPC1-Bsd, plenti-EF1α-IRES-EXTL3-Bsd, pLenti-EF1α-IRES-NPC1ΔA-Bsd, pLenti-EF1α-IRES-NPC1ΔC-Bsd and pLenti-EF1α-IRES-NPC1ΔI-Bsd). At 6 h after transfection, the medium was replaced with fresh medium. The medium supernatant containing LVs was harvested at 48–60 h post-transfection by centrifugation at 2000 rpm (420 × g ) for 10 min, filtrated through a 0.45 μm filter (Merck, Darmstadt, Germany) and stored at −80 °C.
HeLa cells and HEK-293T cells were transduced with LVs in the presence of polybrene (10 μg/mL, Merck) using spinfection through centrifugation at 2000 rpm (420 × g ) for 2 h. At 24 h post-transduction, LV-containing medium was removed and cells were cultured in fresh medium in the presence of 1–2 μg/mL puromycin for 3 to 5 days or 2–5 μg/mL blasticidin (Thermo) for 7–10 days to remove empty cells containing no LVs. Finally, survived cells were collected, aliquoted, and stored in liquid nitrogen.
Baculovirus production and transduction
Bac-EGFP recombinant viruses were generated using Bac-to-Bac baculoviral expression system (Thermo, Cat. No. A11101). Transgene plasmid pFastBac carrying EGFP gene was transformed into DH10Bac E. coli competent cells that harbored the parental bacmid to form a recombinant bacmid encoding the EGFP transgene. The recombinant bacmid was then transfected into insect cells for production of recombinant baculovirus particles. At 72 h post-transfection, the culture supernatant containing baculovirus particles was harvested and concentrated by Optima XPN-100 Ultracentrifuge (Beckman Coulter, California, USA). Virus titer was determined using the 50% tissue culture infectious dose (TCID 50 ) assay 94 .
For baculovirus transduction, HEK-293T or HeLa cells were seeded at a cell density of 40%. At 24 h after seeding, the cells were transduced with baculovirus at an MOI of 1 or 2 for HEK-293T or HeLa respectively. MOIs other than 1 and 2 were indicated in figure legends. At 24 h after baculovirus transduction, baculovirus-containing medium was removed and the cells were incubated in DMEM (Thermo) supplemented with 10% FBS (Thermo) for another 24 h. Thereafter, the cells were collected and the efficiency of baculovirus transduction was quantified by flow cytometry (CytoFLEX, Beckman Coulter, California, USA) and visualized by fluorescence microscope (EVOS M5000, Thermo). The expression of EGFP mRNA was quantified by RT-qPCR using specific primers (Supplementary Table 2 ).
Construction of surfaceome CRISPR library
The human surfaceome CRISPR library was described in our previous work 62 , which contained 16,975 sgRNAs targeting 1314 surface protein genes and 1000 non-targeting sgRNAs. To construct surfaceome CRISPR library in HEK-293T, the cells were transduced with LV library at an MOI of 0.3 using spinfection as described above. Cells of more than 500-fold coverage of the library size were collected, aliquoted, and stored in liquid nitrogen.
CRISPR screening for baculovirus host factors and next-generation sequencing (NGS) analyses of sgRNA enrichment
The HEK-293T cell library of 1 × 10 7 cells was seeded onto 15 cm petri dishes. At 24 h after seeding, the cells were incubated with recombinant baculovirus carrying an EGFP transgene (Bac-EGFP) of 3 × 10 7 infective units (IU) for 24 h. The medium containing virus was then removed and cells washed with phosphate-buffered saline (PBS) for three times and then cultured in fresh medium for another 24 h. Then the treated cells were harvested and sorted using flow cytometry (Moflo, Beckman Coulter, California, USA) for EGFP-negative cells. Genomic DNA of the sorted cells was extracted using phenol: chloroform: isoamyl alcohol (v/v/v, 25:24:1) and then purified using ethanol precipitation. Genome-integrated sgRNAs were amplified from the collected genomic DNA by PCR using primers containing Illumina adapters (Supplementary Table 1 ). PCR amplicons were analyzed by Genewiz (Suzhou, Jiangsu, China) using next-generation sequencing (NGS) on Illumina HiSeq 3000 platform. After removing the adapters, the 20 bp sgRNA was mapped to the reference sgRNA with 1 bp mismatch allowed. The raw read counts were subjected to MAGeCK analyses to determine the enriched sgRNA and gene knockouts. A false discovery rate (FDR) of less than 0.05 was applied to identify significantly enriched sgRNAs and candidate gene knockouts.
Generation of gene knockout cells
The LV transfer plasmids pLentiCRISPR-v2-sgRNA carrying single sgRNAs (Supplementary Table 3 ) for knockout cell line construction were generated as previously described above 62 . The LVs were packaged and transduced onto cells as described above. To evaluate the knockout efficiency, the genomic DNA of knockout cells was extracted using Quick Extraction kit (Lucigen, Wisconsin, USA) and the modified genomic sites were PCR amplified using corresponding primers (Supplementary Table 4 ). The PCR amplicons were sequenced by Sanger sequencing (Genewiz) and gene disruption efficiency was analyzed by TIDE website ( https://tide.nki.nl/ ) 95 . Single clones were obtained by cell sorting using flow cytometry (Moflo, Beckman) and genotyped by Sanger sequencing to determine the mutations at each allele.
Total RNA from cultured cells was extracted using TRIzol (Thermo), chloroform (Sinopharm, Ningbo, China) and purified using isopropanol precipitation. RNA was then reverse transcribed into cDNA by PrimeScript RT reagent Kit with gDNA Eraser (Takara Bio Inc., Shiga, Japan). The mRNA levels were determined by reverse transcription quantitative PCR (RT-qPCR) using SYBR green dye on Applied Biosystems Q6 Real-Time PCR cycler (Thermo) and specific primers (Supplementary Table 2 ). All SYBR Green primers were validated with dissociation curves. The expression of genes was normalized to β-actin or RPLP0 .
For RT-qPCR on mouse tissues, the tissue samples were collected, frozen in liquid nitrogen, and homogenized using grinding beads through tissue grinding device (Jingxin, Shanghai, China) in TRIzol reagent (Thermo). The extraction and quantification processes in tissue RNA were the same to that in cell culture samples as described above.
qPCR quantification of baculovirus genome
Genomic DNA of baculovirus or baculovirus-containing cells was extracted using phenol: chloroform: isoamyl alcohol (v/v/v, 25: 24: 1) and then purified using ethanol precipitation. The total DNA concentration was determined for each sample. For quantitative PCR (qPCR) reaction, 2 ng total DNA was added into each reaction as the template. GP64 DNA, which was used as indicator of viral genome content, was quantified by qPCR using SYBR green dye on Applied Biosystems Q6 Real-Time PCR cycler (Thermo) and specific primers (Supplementary Table 2 ). All SYBR Green primers were validated with dissociation curves. The genomic DNA level was normalized to β-actin.
Gene knockdown using siRNA
HEK-293T cells were seeded onto 6-well plates with a density of 5 × 10 5 cells per well. At 24 h after seeding, cells were transfected with 100 pmol siRNA (Genepharma, Shanghai, China) (Supplementary Table 5 ) using 7.5 μL Lipofectamine 2000 (Thermo) for 6 h, then washed with PBS and cultured in fresh DMEM (Thermo) supplemented with 10% fetal bovine serum (FBS, Thermo). At 48 h post-transfection, cells were infected with baculovirus at an MOI of 1 for 24 h, then washed with PBS for three times and cultured in fresh medium for 24 h. The cell samples were harvested and lysed for total RNA extraction, and the mRNA levels of EGFP , EXTL3 , and NPC1 in cell lysate were determined using RT-qPCR as described above.
Virus attachment and entry assays
Virus attachment and entry assays were performed as described 96 with minor modifications. HEK-293T cells or HeLa cells were seeded onto 12-well plates at a density of 200,000 cells per well and incubated overnight. For virus attachment assay, cells were incubated with baculovirus at an MOI of 20 in cold medium without FBS on ice for 60 min, then washed with cold Dulbecco’s phosphate-buffered saline (DPBS) for three times and harvested. The baculovirus genomic DNA was extracted from the cells containing attached virus and quantified by qPCR as described above. For virus entry assay, cells were incubated with baculovirus at an MOI of 20 in cold medium on ice for 60 min, washed by cold DPBS for three times, treated with pre-warmed medium containing FBS, and then incubated at 37 °C for 40 min. The treated cells were washed with PBS for three times and then treated with 0.25% trypsin (Thermo) for 30 s to remove surface-bound baculovirus particles. The genomic DNA of internalized baculovirus was extracted and quantified by qPCR as described above.
Construction of stable cell lines harboring overexpressed genes
EXTL3 and NPC1 genes were codon-optimized for expression in human cells and synthesized by Genewiz (Supplementary Tables 6 , 7 ). The 20 bp sgRNA-targeting sites and PAM sequences were mutated with silent mutations. Myc and FLAG tags were added to the C-terminus of these genes for WB detection. These genes were cloned into the XbaI and BamH1 sites of pLV-EF1α-IRES-Bsd plasmid. The lentiviral overexpression plasmids of NPC1 truncation mutants (NPC1-ΔA, NPC1-ΔC, and NPC1-ΔI) were constructed from the full-length gene in pLV-EF1ɑ-IRES-NPC1-Bsd. For LV packaging, the transgene plasmid containing EXTL3, NPC1, NPC1ΔA, NPC1ΔC or NPC1ΔI was co-transfected into HEK-293T cells with helper plasmids pMD2.G and psPAX as described above. The transgene-containing LVs were transduced into EXTL3 −/− or NPC1 −/− cells, and the cells were subjected to selection with 2 μg/mL blasticidin (Thermo) for 7 to 10 days to purge empty cells containing no LVs. Finally, survived cells were collected, aliquoted, and stored in liquid nitrogen.
Analysis of the intracellular baculovirus genomic DNA
The experiment was performed in an approach similar to a previous study 62 . Non-targeting sgRNA-treated, NPC1 −/− and NPC1 overexpression rescued HeLa cells were seeded onto 24-well plates at a density of 80,000 cells per well and incubated overnight. The next day the cells were incubated with Bac-EGFP at an MOI of 2 for 1 h. The virus-containing medium was removed and the cells were washed with PBS for three times and then treated with 0.25% trypsin (Thermo) for 30 s to remove surface-bound baculovirus particles. These treated cells were supplemented with pre-warmed DMEM medium containing FBS and then incubated at 37 °C for indicated time. Intracellular baculovirus genomic DNA was extracted from each well. The total baculovirus GP64 amount in each well was determined by qPCR and then normalized to the zero time point.
Cell Counting Kit-8 assay (CCK-8)
Non-targeting sgRNA, NPC1 −/− , and NPC1 rescued HeLa cells were seeded onto 96-well plates at a density of 10,000 cells per well and incubated at 37 °C overnight. The cells were incubated with Bac-EGFP at an MOI of 2 for 1 h, washed with PBS for three times and then treated with 0.25% trypsin (Thermo) for 30 s to remove surface-bound baculovirus particles. At indicated time points, the supernatant was removed and cells incubated with complete medium containing 10% CCK-8 reagent (MeilunBio, Liaoning, China) at 37 °C for 1 h. Cell proliferation was quantified by measuring absorbance at 450 nm using microplate reader (SpectraMax iD3, Molecular Devices, Shanghai, China). The background signal in empty wells without cells was subtracted from each sample.
Bafilomycin A1 (BafA1) treatment
Wild-type HeLa cells were seeded on to 24-well plates at a density of 80,000 cells per well and incubated overnight. The next day the cells were pre-incubated with 100 nM BafA1 (MedChemExpress, New Jersey, USA, Cat. No. HY-100558) for 1 h, followed by transduction with Bac-EGFP at an MOI of 2 for 1 h in the presence of 100 nM BafA1. The virus-containing medium was then removed, the cells washed three times with PBS and then treated with 0.25% trypsin (Thermo) for 30 s to remove surface-bound baculovirus particles. These cells were incubated in DMEM medium containing FBS and 10 nM BafA1 for indicated time. The genomic DNA of intracellular baculovirus was extracted from cells and quantified by qPCR as described above.
For CCK-8 assay, cells were seeded on to 96-well plates at a density of 10,000 cells per well. The next day, the cells were pre-incubated with 100 nM BafA1 and transduced with Bac-EGFP to mimic the treatment as described above. At 48 h after removal of virus, cell proliferation was quantified by CCK-8 as described above at indicated time points.
WB analyses
For WB analyses, cells were lysed with RIPA buffer (Beyotime Biotechnology, Beijing, China) on ice for 10 min and centrifuged at 12,000 rpm (13,523 × g ) at 4 °C for 10 min to remove cell debris. The total protein concentration in cell lysate was determined using the BCA Protein Assay Kit (Thermo). Cell lysate was mixed with SDS-PAGE loading buffer (Takara) containing 200 mM dithiothreitol (DTT), incubated at 95 °C for 10 min, and resolved on 4–12% PAGE gels (GenScript, Nanjing, China). Protein samples were transferred onto nitrocellulose membranes (Merck) using an iBlot gel transfer system (Thermo). The following primary and secondary antibodies were used in WB including anti-Myc rabbit antibody (CST, Cat. No. 2272S), anti-HA rabbit antibody (CST, Cat. No. 3724S), anti-Heparan Sulfate Proteoglycan 2/Perlecan antibody (Abcam, Cambridge, UK, Cat. No. ab255829) and HRP-conjugated anti-rabbit IgG (CST, Cat. No. 5127S). Anti-β actin antibody conjugated with HRP (CST, Cat. No. 5125S) was used as an internal control.
Co-immunoprecipitation (Co-IP)
HEK-293T cells were seeded onto 6-well plates at a density of 1 × 10 6 cells per well. After the confluency reached 70% to 80%, plasmid was transfected into cells. Plasmids containing Myc-labeled NPC1 or NPC1 mutant and plasmids containing HA-labeled GP64 (Supplementary Table 8 ) were co-transfected into HEK-293T cells by Lipofectamine 3000 (Thermo). At 24 h after transfection, cells were resuspended with co-IP lysis buffer. One-half of the lysate was used to incubate with magnetic beads (Thermo) containing anti-HA or anti-Myc antibodies at room temperature for 1 h. Magnetic beads were collected with a magnetic rack (Thermo), then proteins bound to the magnetic beads were eluted by 1x SDS-PAGE loading buffer containing DTT. Finally, total proteins and co-immunoprecipitated proteins were detected by WB as described above.
Recombinant NPC1 protein neutralization assay
HEK-293T cells were seeded on to 48-well plates at a density of 1 × 10 5 cells per well. After Bac-EGFP incubation with recombinant protein of NPC1 C domain (residues R372-F622) (SinoBiological, Beijing, China, Cat. No. 16499-H32H) at indicated concentrations at 4 °C for 30 min. Thereafter, HEK-293T cells were transduced with Bac-EGFP at an MOI of 1 for 24 h in the presence of NPC1-C domain protein. Then baculovirus-containing medium was removed and the cells were incubated in DMEM (Thermo) supplemented with 10% FBS (Thermo) for 24 h and then harvested. The efficiency of baculovirus transduction was quantified by flow cytometry (CytoFLEX, Beckman Coulter, California, USA).
Expression and purification of the human NPC1-I domain
NPC1-I domain was codon-optimized for expression in Escherichia coli, synthesized by Genewiz, and cloned into pGEX4T-1-GST plasmid. The recombinant pGEX4T-1-GST-NPC1-I plasmid was transformed into chemically competent BL21 (DE3) E. coli cells. After overnight culturing, single colonies were selected and amplified in Luria–Bertani (LB) medium containing (100 mg/mL) ampicillin overnight. The next day, the culture was inoculated into fresh LB medium at 1: 100 dilution and cultured at 37 °C for about 3 h until OD 600 reached 0.6–0.8. Protein expression was induced with 0.5 mM isopropyl β-D-thiogalactoside (IPTG) at 22 °C overnight. The cells were then collected by centrifugation at 4,000 rpm (1681 × g ) for 15 min at 4 °C.
For protein purification, the cell pellet was resuspended with PBS and then lysed by sonication. The cell lysate was centrifuged at 12,000 rpm (13,523 × g ) for 30 min at 4 °C and the supernatant was isolated. The supernatant was run through a glutathione resin affinity chromatography column (Merck) for protein capture. The resin was sufficiently washed with PBS and the protein was eluted with wash buffer supplemented with reduced glutathione. The eluted proteins were concentrated and further purified by size exclusion chromatography (Superdex 200 Increase 10/300GL) with a buffer containing 20 mM Tris pH 8.0, 250 mM NaCl. The fractions containing target proteins were pooled and concentrated for mass spectrometry analysis.
Mass spectrometry validation of purified NPC1-I protein
The purified NPC1-I protein was validated by Orbitrap Fusion MS (Thermo Fisher, San Jose, CA) using Proteome Discoverer 2.2 and Xcalibur analysis software at the Analytical Chemistry Platform at SIAIS, ShanghaiTech University. The entire gels were rinsed with water for 3–4 h, then the bands of GST (control, n = 1) and NPC-I (sample, n = 1) were excised with a clean scalpel. The two excised bands were cut into cubes (1 mm × 1 mm) separately. The gel slices were transferred into two microcentrifuge tubes and spun down by a microcentrifuge. For in-gel reduction, alkylation, and de-staining, the gels were incubated with 100 μL de-staining solution (100 mM ammonium bicarbonate buffer containing 50% acetonitrile, vol/vol) for 30 min at 37 °C, and then incubated with 500 μL neat acetonitrile for 10 min until gel slices shrank and became opaque and stiff. These gel slices were span to remove all liquid and 40 μL of 10 mM DTT solution was added to cover gel slices for 30 min at 56 °C in an air thermostat. The tubes were chilled down to room temperature, and incubated with 500 μL acetonitrile for 10 min and then all liquid was removed. The gels were incubated with 40 μL of 55 mM iodoacetamide solution for 20 min at room temperature in the dark, treated with acetonitrile, and then all liquid removed. The dry gels were incubated with 70 μL trypsin buffer containing 13 μg/mL trypsin for 100 min at 4 °C and then incubated with 20 μL 100 mM ammonium bicarbonate containing 10% (vol/vol) acetonitrile at 37 °C overnight. The next day the samples were incubated with 200 μL extraction buffer (1 vol: 2 vol of 5% formic acid: acetonitrile) for 15 min at 37 °C in a shaker. Then the supernatant was collected into a PCR tube, dried in a vacuum centrifuge, and stored at 4 °C for further analysis.
For liquid chromatography (LC) analysis, the above samples were supplemented with 20 μL of 0.1% (vol/vol) formic acid, vortexed, and centrifuged for 5 min at 12,000 × g . These treated peptides were loaded onto an analytical column (Ionopticks, AUR2-25075C18A, 25 cm × 75 μm, C18, 1.6 μm) connected to an Easy-nLC1200 UHPLC-Orbitrap Fusion (Thermo Fisher Scientific). The elution gradient and mobile phase constitution used for peptide separation were set as follows: 0–60 min, 5–30% buffer B; 60–70 min, 30–45% buffer B; 70–75 min, 45–100% buffer B; 75–80 min, 100% buffer B at a flow rate of 300 nL/min. Mobile phase in buffer A was 0.1% formic acid in water and mobile phase in buffer B was 0.1% formic acid in 80% acetonitrile.
Peptides eluted from the LC column were directly electro-sprayed into the mass spectrometer with the application of a distal 2.1 kV spray voltage. Survey full-scan mass spectra (from m/z 350 to 1800) were acquired in the Orbitrap analyzer (Orbitrap Fusion, Thermo Fisher Scientific) with resolution r = 60,000 at m/z 400. The cycle time of the MS-MS2 events was 3 s, sequentially generated and selected from the full mass spectrum at a 32% normalized collision energy. The dynamic exclusion time was set to 10 s. The acquired MS/MS data were analyzed using the AA sequence of protein GST, using Protein Discoverer 2.2 with the parameter settings as follows: precursor and fragment mass tolerance of 10 ppm and 0.02 Da and dynamic modifications of + 15.995 Da for Oxidation (Met) and + 42.011 Da for Acetyl (N-terminus), and + 57.021 Da for Carbamidomethyl (C terminus) as static modifications. To accurately estimate peptide probabilities and filter the false discovery, the fixed value PSM validator node was used and set to have a maximum delta Cn of 0.05. Trypsin was defined as cleavage enzyme and the maximal number of missed cleavage sites was set to two.
Far western blotting
Far western blotting was performed as previously described 97 . Briefly, 0.2–1 μg purified NPC1-I protein (with an unknown impurity band) or a control GST protein was mixed with SDS-PAGE loading buffer (Takara) containing 200 mM DTT, incubated at 95 °C for 10 min and resolved on 4–12% PAGE gels (GenScript, Nanjing, China). Protein samples were transferred onto nitrocellulose (PVDF) membranes (Merck) using an iBlot gel transfer system (Thermo). PVDF membrane was treated with PBS solution containing gradient concentrations of guanidine hydrochloride (GuHCl) solution (Solarbio, China) for protein unfolding and refolding. The GuHCl gradient was decreased from 6 M to 3 M and then 0.1 M with the membrane being treated with each gradient for 30 min. Finally, the membrane was treated with PBS without GuHCl at 4 °C overnight. The membrane was then blocked with PBS containing 5% milk (w/v) and incubated with 1–10 μg GP64 protein (SinoBiological, Beijing, China) overnight at 4 °C. The membrane was then treated with an anti-GP64 mouse antibody (Abcam, Cat. No. ab91214) as the primary antibody and an HRP-conjugated anti-mouse antibody (R&D, Cat. No. HAF007) as the secondary antibody. The bait protein could be detected at the location of the prey protein on the membrane if they interact to form a complex.
Transduction with DiOC18-labeled baculovirus
Baculovirus was labeled with DiOC18 (Thermo) at a final concentration of 2 μM in PBS. The solution was rotated at room temperature in the dark for 1 h and then filtered through a 0.22 μm pore size syringe filter (Merck) to remove unbound dyes and aggregates. Cells were infected with DiOC18-labeled baculovirus at 37 °C for 3 h, then washed with PBS three times and treated with trypsin briefly to remove free and surface-bound viruses. Thereafter, the cells were fixed with PFA and analyzed by flow cytometry (CytoFLEX, Beckman Coulter, California, USA) and confocal microscopy (LSM 710, Zeiss, Oberkochen, Germany). A control group with DiOC18 dye alone, without the addition of baculovirus, was prepared following the same filtration process as described above to assess the background fluorescence labeling of cells by free dyes in the DiOC18-Bac-EGFP-treated groups.
Baculovirus transduction in mice
Npc1 −/+ and wild-type C57BL/6 mice were purchased from, bred, and raised in GemPharmatech (Nanjing, China). All mice were housed in animal facility with ambient room temperature of 20–26 °C, humidity of 50–70%, and dark/light cycle of 12 h. Baculovirus-related animal experiments were performed in OBiO-tech corporation (Shanghai, China). For baculovirus, transduction, 6–8-week-old male Npc1 −/+ and wild-type C57BL/6 mice were used. The mice were maintained within a Specific Pathogen-Free (SPF) facility with free access to water and food. The housing facility for mice was under a 12:12 h light: dark cycle at temperatures 20–26 °C, humidity 40–70%. For transduction experiments, mice were randomly grouped ( n = 3 to 9 per group as indicated) and injected with baculoviruses through tail vein with a dose of 4.4 × 10 4 IU per gram body weight. Animals were sacrificed under an anesthetic condition at the indicated time post-baculovirus transduction. The tissues including liver, brain, spleen, lung, and kidney were collected and flash-frozen in liquid nitrogen or fixed in PFA buffer for further analyses.
For tissue tropism of Bac-EGFP transduction, Bac-EGFP-transduced tissues were flash-frozen and fixed with optimal cutting temperature compound (OCT). The fixed tissues were sectioned at 5 μm, and the sections were fixed with PFA and stained with Hoechst (Thermo). The fluorescence images were acquired using confocal microscopy (LSM 710, Zeiss).
Statistics and reproducibility
All data were the results from at least three biological replicates and were shown as mean ± standard deviation unless noted otherwise. All experiments were repeated three times independently (biological replicates) unless noted otherwise. No data were excluded for analyses. Statistical analyses and graphing were performed with GraphPad Prism 7.0. The P -values were determined using two-tailed unpaired Student’s t-test unless otherwise noted.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that all data supporting the findings of this study are available in the article, its Supplementary Data, its Source Data, or from the corresponding authors upon request. Source data are provided with this paper. The NGS data generated in this study have been deposited in the SRA database under accession code PRJNA1131913 . Source data are provided with this paper.
Ihalainen, T. O. et al. Morphological characterization of baculovirus Autographa californica multiple nucleopolyhedrovirus. Virus Res. 148 , 71–74 (2010).
Article CAS PubMed Google Scholar
Ayres, M. D., Howard, S. C., Kuzio, J., Lopez-Ferber, M. & Possee, R. D. The complete DNA sequence of Autographa californica nuclear polyhedrosis virus. Virology 202 , 586–605 (1994).
Rohrmann, G. F. Baculovirus structural proteins. J. Gen. Virol. 73 , 749–761 (1992).
Summers, M. D. & Volkman, L. E. Comparison of biophysical and morphological properties of occluded and extracellular nonoccluded baculovirus from in vivo and in vitro host systems. J. Virol. 17 , 962–972 (1976).
Article CAS PubMed PubMed Central Google Scholar
Blissard, G. W. & Rohrmann, G. F. Location, sequence, transcriptional mapping, and temporal expression of the gp64 envelope glycoprotein gene of the Orgyia pseudotsugata multicapsid nuclear polyhedrosis virus. Virology 170 , 537–555 (1989).
Peng, K. et al. Characterization of novel components of the baculovirus per os infectivity factor complex. J. Virol. 86 , 4981–4988 (2012).
Peng, K., van Oers, M. M., Hu, Z., van Lent, J. W. & Vlak, J. M. Baculovirus per os infectivity factors form a complex on the surface of occlusion-derived virus. J. Virol. 84 , 9497–9504 (2010).
Volkman, L. E. & Summers, M. D. Autographa californica nuclear polyhedrosis virus: comparative infectivity of the occluded, alkali-liberated, and nonoccluded forms. J. Invertebr. Pathol. 30 , 102–103 (1977).
Summers, M. D. Electron microscopic observations on granulosis virus entry, uncoating and replication processes during infection of the midgut cells of Trichoplusia ni. J. Ultrastruct. 35 , 606–625 (1971).
Article CAS Google Scholar
Zhou, J. & Blissard, G. W. Identification of a GP64 subdomain involved in receptor binding by budded virions of the baculovirus Autographica californica multicapsid nucleopolyhedrovirus. J. Virol. 82 , 4449–4460 (2008).
Long, G., Pan, X., Kormelink, R. & Vlak, J. M. Functional entry of baculovirus into insect and mammalian cells is dependent on clathrin-mediated endocytosis. J. Virol. 80 , 8830–8833 (2006).
Granados, R. R. & Lawler, K. A. In vivo pathway of Autographa californica baculovirus invasion and infection. Virology 108 , 297–308 (1981).
Horton, H. M. & Burand, J. P. Saturable attachment sites for polyhedron-derived baculovirus on insect cells and evidence for entry via direct membrane fusion. J. Virol. 67 , 1860–1868 (1993).
Pearson, M. N., Russell, R. L. & Rohrmann, G. F. Characterization of a baculovirus-encoded protein that is associated with infected-cell membranes and budded virions. Virology 291 , 22–31 (2001).
Lung, O. Y., Cruz-Alvarez, M. & Blissard, G. W. Ac23, an envelope fusion protein homolog in the baculovirus Autographa californica multicapsid nucleopolyhedrovirus, is a viral pathogenicity factor. J. Virol. 77 , 328–339 (2003).
WF, I. J. et al. A novel baculovirus envelope fusion protein with a proprotein convertase cleavage site. Virology 275 , 30–41 (2000).
Article Google Scholar
Long, G., Pan, X., Westenberg, M. & Vlak, J. M. Functional role of the cytoplasmic tail domain of the major envelope fusion protein of group II baculoviruses. J. Virol. 80 , 11226–11234 (2006).
Westenberg, M. et al. Functional analysis of the putative fusion domain of the baculovirus envelope fusion protein F. J. Virol. 78 , 6946–6954 (2004).
Kadlec, J., Loureiro, S., Abrescia, N. G., Stuart, D. I. & Jones, I. M. The postfusion structure of baculovirus gp64 supports a unified view of viral fusion machines. Nat. Struct. Mol. Biol. 15 , 1024–1030 (2008).
White, J. M., Delos, S. E., Brecher, M. & Schornberg, K. Structures and mechanisms of viral membrane fusion proteins: multiple variations on a common theme. Crit. Rev. Biochem. 43 , 189–219 (2008).
Blissard, G. W. & Wenz, J. R. Baculovirus gp64 envelope glycoprotein is sufficient to mediate pH-dependent membrane fusion. J. Virol. 66 , 6829–6835 (1992).
Zhou, J. & Blissard, G. W. Mapping the conformational epitope of a neutralizing antibody (AcV1) directed against the AcMNPV GP64 protein. Virology 352 , 427–437 (2006).
Rohrmann, G. F. The AcMNPV genome: gene content, conservation, and function. in Baculovirus Molecular Biology , 3rd edn, 1–48 (National Center for Biotechnology Information. Bethesda, MD, 2013).
Smith, G. E., Summers, M. D. & Fraser, M. J. Production of human beta interferon in insect cells infected with a baculovirus expression vector. MCB 3 , 2156–2165 (1983).
CAS PubMed PubMed Central Google Scholar
Potter, K. N., Li, Y. & Capra, J. D. Antibody production in the baculovirus expression system. Int Rev. Immunol. 10 , 103–112 (1993).
Gheysen, D. et al. Assembly and release of HIV-1 precursor Pr55gag virus-like particles from recombinant baculovirus-infected insect cells. Cell 59 , 103–112 (1989).
Malogolovkin, A. et al. Enhanced Zika virus-like particle development using Baculovirus spp. constructs. J. Med. Virol. 95 , e28252 (2023).
Kim, C. H. et al. Direct vaccination with pseudotype baculovirus expressing murine telomerase induces anti-tumor immunity comparable with RNA-electroporated dendritic cells in a murine glioma model. Cancer Lett. 250 , 276–283 (2007).
Fan, H. et al. Construction and immunogenicity of recombinant pseudotype baculovirus expressing the capsid protein of porcine circovirus type 2 in mice. J. Virol. Methods 150 , 21–26 (2008).
Huang, H. et al. Construction and immunogenicity of a recombinant pseudotype baculovirus expressing the glycoprotein of rabies virus in mice. Arch. Virol. 156 , 753–758 (2011).
Wang, S. et al. Construction and immunogenicity of pseudotype baculovirus expressing GP5 and M protein of porcine reproductive and respiratory syndrome virus. Vaccine 25 , 8220–8227 (2007).
Fang, R. et al. Construction and immunogenicity of pseudotype baculovirus expressing Toxoplasma gondii SAG1 protein in BALB/c mice model. Vaccine 28 , 1803–1807 (2010).
Grabowska, A. K. et al. New baculovirus recombinants expressing Pseudorabies virus (PRV) glycoproteins protect mice against lethal challenge infection. Vaccine 27 , 3584–3591 (2009).
Chen, C. Y., Lin, C. Y., Chen, G. Y. & Hu, Y. C. Baculovirus as a gene delivery vector: recent understandings of molecular alterations in transduced cells and latest applications. Biotechnol. Adv. 29 , 618–631 (2011).
Makkonen, K. E., Airenne, K. & Ylä-Herttulala, S. Baculovirus-mediated gene delivery and RNAi applications. Viruses 7 , 2099–2125 (2015).
Mansouri, M. & Berger, P. Baculovirus for gene delivery to mammalian cells: Past, present and future. Plasmid 98 , 1–7 (2018).
Lei, Y. et al. Gene editing of human embryonic stem cells via an engineered baculoviral vector carrying zinc-finger nucleases. Mol. Ther. 19 , 942–950 (2011).
Tay, F. C. et al. Targeted transgene insertion into the AAVS1 locus driven by baculoviral vector-mediated zinc finger nuclease expression in human-induced pluripotent stem cells. J. Gene Med. 15 , 384–395 (2013).
Phang, R. Z. et al. Zinc finger nuclease-expressing baculoviral vectors mediate targeted genome integration of reprogramming factor genes to facilitate the generation of human induced pluripotent stem cells. Stem Cells Transl. Med. 2 , 935–945 (2013).
Zhu, H. et al. Baculoviral transduction facilitates TALEN-mediated targeted transgene integration and Cre/LoxP cassette exchange in human-induced pluripotent stem cells. Nucleic Acids Res. 41 , e180 (2013).
Hindriksen, S. et al. Baculoviral delivery of CRISPR/Cas9 facilitates efficient genome editing in human cells. PLoS One 12 , e0179514 (2017).
Article PubMed PubMed Central Google Scholar
Aulicino, F. et al. Highly efficient CRISPR-mediated large DNA docking and multiplexed prime editing using a single baculovirus. Nucleic Acids Res. 50 , 7783–7799 (2022).
Nguyen, G. N. et al. A long-term study of AAV gene therapy in dogs with hemophilia A identifies clonal expansions of transduced liver cells. Nat. Biotechnol. 39 , 47–55 (2021).
McCarty, D. M., Young, S. M. Jr. & Samulski, R. J. Integration of adeno-associated virus (AAV) and recombinant AAV vectors. Annu Rev. Genet. 38 , 819–845 (2004).
Sarkis, C. et al. Efficient transduction of neural cells in vitro and in vivo by a baculovirus-derived vector. Proc. Natl Acad. Sci. USA 97 , 14638–14643 (2000).
Article ADS CAS PubMed PubMed Central Google Scholar
Turunen, T. A., Laakkonen, J. P., Alasaarela, L., Airenne, K. J. & Ylä-Herttuala, S. Sleeping Beauty-baculovirus hybrid vectors for long-term gene expression in the eye. J. Gene Med. 16 , 40–53 (2014).
Kinnunen, K. et al. Baculovirus is an efficient vector for the transduction of the eye: comparison of baculovirus- and adenovirus-mediated intravitreal vascular endothelial growth factor D gene transfer in the rabbit eye. J. Gene Med. 11 , 382–389 (2009).
Luo, W. Y. et al. Baculovirus vectors for antiangiogenesis-based cancer gene therapy. Cancer Gene Ther. 18 , 637–645 (2011).
Luo, W. Y. et al. Development of the hybrid Sleeping Beauty: baculovirus vector for sustained gene expression and cancer therapy. Gene Ther. 19 , 844–851 (2012).
Liu, Z. et al. Surface displaying of swine IgG1 Fc enhances baculovirus-vectored vaccine efficacy by facilitating viral complement escape and mammalian cell transduction. Vet. Res. 48 , 29 (2017).
Article ADS PubMed PubMed Central Google Scholar
Hu, L. et al. Improving baculovirus transduction of mammalian cells by incorporation of thogotovirus glycoproteins. Virol. Sin. 34 , 454–466 (2019).
Article MathSciNet PubMed PubMed Central Google Scholar
Pieroni, L., Maione, D. & La Monica, N. In vivo gene transfer in mouse skeletal muscle mediated by baculovirus vectors. Hum. Gene Ther. 12 , 871–881 (2001).
Barsoum, J., Brown, R., McKee, M. & Boyce, F. M. Efficient transduction of mammalian cells by a recombinant baculovirus having the vesicular stomatitis virus G glycoprotein. Hum. Gene Ther. 8 , 2011–2018 (1997).
Duisit, G. et al. Baculovirus vector requires electrostatic interactions including heparan sulfate for efficient gene transfer in mammalian cells. J. Gene Med. 1 , 93–102 (1999).
Wu, C. & Wang, S. A pH-sensitive heparin-binding sequence from Baculovirus gp64 protein is important for binding to mammalian cells but not to Sf9 insect cells. J. Virol. 86 , 484–491 (2012).
Makkonen, K. E. et al. 6-o- and N-sulfated syndecan-1 promotes baculovirus binding and entry into Mammalian cells. J. Virol. 87 , 11148–11159 (2013).
Tani, H., Nishijima, M., Ushijima, H., Miyamura, T. & Matsuura, Y. Characterization of cell-surface determinants important for baculovirus infection. Virology 279 , 343–353 (2001).
Luz-Madrigal, A., Asanov, A., Camacho-Zarco, A. R., Sampieri, A. & Vaca, L. A cholesterol recognition amino acid consensus domain in GP64 fusion protein facilitates anchoring of baculovirus to mammalian cells. J. Virol. 87 , 11894–11907 (2013).
Kataoka, C. et al. Baculovirus GP64-mediated entry into mammalian cells. J. Virol. 86 , 2610–2620 (2012).
Amalfi, S. et al. Baculovirus transduction in mammalian cells is affected by the production of type I and III interferons, which is mediated mainly by the cGAS-STING pathway. J. Virol. 94 , e01555-20 (2020).
Puschnik, A. S., Majzoub, K., Ooi, Y. S. & Carette, J. E. A CRISPR toolbox to study virus-host interactions. Nat. Rev. Microbiol. 15 , 351–364 (2017).
Mei, H. et al. Surfaceome CRISPR screen identifies OLFML3 as a rhinovirus-inducible IFN antagonist. Genome Biol. 22 , 297 (2021).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343 , 84–87 (2014).
Article ADS CAS PubMed Google Scholar
Clausen, T. M. et al. SARS-CoV-2 infection depends on cellular heparan sulfate and ACE2. Cell 183 , 1043–1057.e1015 (2020).
Shukla, D. et al. A novel role for 3-O-sulfated heparan sulfate in herpes simplex virus 1 entry. Cell 99 , 13–22 (1999).
Chen, Y. et al. Dengue virus infectivity depends on envelope protein binding to target cell heparan sulfate. Nat. Med. 3 , 866–871 (1997).
Carette, J. E. et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature 477 , 340–343 (2011).
King, L. B. et al. The Marburgvirus-neutralizing human monoclonal antibody MR191 targets a conserved site to block virus receptor binding. Cell Host Microbe 23 , 101–109.e104 (2018).
Tjia, S. T., zu Altenschildesche, G. M. & Doerfler, W. Autographa californica nuclear polyhedrosis virus (AcNPV) DNA does not persist in mass cultures of mammalian cells. Virology 125 , 107–117 (1983).
Bowman, E. J., Siebers, A. & Altendorf, K. Bafilomycins: a class of inhibitors of membrane ATPases from microorganisms, animal cells, and plant cells. Proc. Natl Acad. Sci. USA 85 , 7972–7976 (1988).
Zhao, H. et al. A broad-spectrum virus- and host-targeting peptide against respiratory viruses including influenza virus and SARS-CoV-2. Nat. Commun. 11 , 4252 (2020).
Shang, C. et al. Inhibitors of endosomal acidification suppress SARS-CoV-2 replication and relieve viral pneumonia in hACE2 transgenic mice. Virol. J. 18 , 46 (2021).
Yamada, S. Specific functions of Exostosin-like 3 (EXTL3) gene products. Cell Mol. Biol. Lett. 25 , 39 (2020).
Awad, W., Kjellström, S., Svensson Birkedal, G., Mani, K. & Logan, D. T. Structural and biophysical characterization of human EXTL3: domain organization, glycosylation, and solution structure. Biochemistry 57 , 1166–1177 (2018).
Dogra, P. et al. Novel heparan sulfate-binding peptides for blocking herpesvirus entry. PLoS One 10 , e0126239 (2015).
Sarrazin, S., Lamanna, W. C. & Esko, J. D. Heparan sulfate proteoglycans. Cold Spring Harb. Perspect. Biol. 3 , a004952 (2011).
Giraldo, M. I. et al. Envelope protein ubiquitination drives entry and pathogenesis of Zika virus. Nature 585 , 414–419 (2020).
Banerjee, I. et al. Influenza A virus uses the aggresome processing machinery for host cell entry. Science 346 , 473–477 (2014).
Davies, J. P., Chen, F. W. & Ioannou, Y. A. Transmembrane molecular pump activity of Niemann-Pick C1 protein. Science 290 , 2295–2298 (2000).
Wang, H. et al. Ebola viral glycoprotein bound to its endosomal receptor Niemann-Pick C1. Cell 164 , 258–268 (2016).
Gong, X. et al. Structural insights into the Niemann-Pick C1 (NPC1)-mediated cholesterol transfer and Ebola infection. Cell 165 , 1467–1478 (2016).
Santiago-Mujica, E. et al. Hepatic and neuronal phenotype of NPC1(−/−) mice. Heliyon 5 , e01293 (2019).
Berger, I., Fitzgerald, D. J. & Richmond, T. J. Baculovirus expression system for heterologous multiprotein complexes. Nat. Biotechnol. 22 , 1583–1587 (2004).
Fitzgerald, D. J. et al. Protein complex expression by using multigene baculoviral vectors. Nat. Methods 3 , 1021–1032 (2006).
Bieniossek, C. et al. Automated unrestricted multigene recombineering for multiprotein complex production. Nat. Methods 6 , 447–450 (2009).
Hunt, C. L., Lennemann, N. J. & Maury, W. Filovirus entry: a novelty in the viral fusion world. Viruses 4 , 258–275 (2012).
Cuesta-Geijo, M. et al. New insights into the role of endosomal proteins for African swine fever virus infection. Plos Pathog. 18 , e1009784 (2022).
Fan, Y. et al. The NPC families mediate BmNPV entry. Microbiol. Spectr. 10 , e0091722 (2022).
Article PubMed Google Scholar
Li, Z. et al. Baculovirus utilizes cholesterol transporter Niemann-Pick C1 for host cell entry. Front. Microbiol. 10 , 2825 (2019).
Ye, J., Davé, U. P., Grishin, N. V., Goldstein, J. L. & Brown, M. S. Asparagine-proline sequence within membrane-spanning segment of SREBP triggers intramembrane cleavage by site-2 protease. Proc. Natl Acad. Sci. USA 97 , 5123–5128 (2000).
Ye, J. et al. ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs. Mol. Cell 6 , 1355–1364 (2000).
Denard, B. et al. The membrane-bound transcription factor CREB3L1 is activated in response to virus infection to inhibit proliferation of virus-infected cells. Cell Host Microbe 10 , 65–74 (2011).
Amalfi, S. et al. P26 enhances baculovirus gene delivery by modulating the mammalian antiviral response. Appl. Microbiol Biot. 107 , 6277–6286 (2023).
Reed, L. J. & Muench, H. A Simple method of estimating fifty percent endpoints. Am. J. Epidemiol. 27 , 493–497 (1938).
Brinkman, E. K. et al. Easy quantification of template-directed CRISPR/Cas9 editing. Nucleic Acids Res. 46 , e58 (2018).
Xu, Z. S. et al. LDLR is an entry receptor for Crimean-Congo hemorrhagic fever virus. Cell Res. 34 , 140–150 (2024).
Wu, Y., Li, Q. & Chen, X. Z. Detecting protein-protein interactions by Far western blotting. Nat. Protoc. 2 , 3278–3284 (2007).
Download references
Acknowledgements
We thank Lishuang Zhang and Pengwei Zhang from the Discovery Technology Platform of SIAIS for the support of flow cytometry experiments, Wei Zhu from the Analytical Chemistry Platform for assistance with mass spectrometry analysis, and the Biomedical Big Data Platform for the design of sgRNA libraries and MAGeCK analyses. This research was supported by the Science and Technology Commission of Shanghai Municipality (23ZR1442100 to J.L.), the Shanghai Frontiers Science Center for Biomacromolecules and Precision Medicine at ShanghaiTech University (2022A0301-417-01 to J.L.), and the Postgraduate Scientific Research Innovation Project of Central South University (2023ZZTS0542 to Y.H.).
Author information
These authors contributed equally: Yuege Huang, Hong Mei.
Authors and Affiliations
Furong Laboratory, Center for Medical Genetics, School of Life Sciences, Central South University, Changsha, Hunan, China
Yuege Huang & Jia-Da Li
Shanghai Institute for Advanced Immunochemical Studies, ShanghaiTech University, Shanghai, China
Hong Mei, Chunchen Deng, Wei Wang, Chao Yuan, Yan Nie & Jia Liu
School of Life Science and Technology, ShanghaiTech University, Shanghai, China
Chunchen Deng, Chao Yuan & Jia Liu
Hunan Key Laboratory of Animal Models for Human Diseases, Changsha, Hunan, China
Shanghai Clinical Research and Trial Center, Shanghai, China
Guangzhou Laboratory, Guangzhou International Bio Island, Guangzhou, Guangdong, China
Shanghai Asiflyerbio Biotechnology, Shanghai, China
You can also search for this author in PubMed Google Scholar
Contributions
J.L., J.D.L., and H.M. conceived this study, J.L., H.M., Y.H., and Y.N. designed the experiments, and Y.H., H.M., C.D., and C.Y. carried out the experiments and analyzed the data. W.W., H.M., and L.J. designed the sgRNA libraries, and Y.H., H.M., J.L, and J.D.L. wrote the manuscript. All authors contributed to the manuscript and approved the submission.
Corresponding authors
Correspondence to Hong Mei , Jia-Da Li or Jia Liu .
Ethics declarations
Competing interests.
J. Liu is the founder and shareholder of Shanghai AsiFlyer Biotechnology. The remaining authors declare no competing interests.
Ethical approval
All study protocols involving mice were approved by the ethical committee of ShanghaiTech University (approval number: 20230821001) and conducted in accordance with regulatory policies in China for the care and use of animals.
Peer review
Peer review information.
Nature Communications thanks Francesco Aulicino and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information, peer review file, description of additional supplementary files, supplementary data 1, reporting summary, source data, source data, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/ .
Reprints and permissions
About this article
Cite this article.
Huang, Y., Mei, H., Deng, C. et al. EXTL3 and NPC1 are mammalian host factors for Autographa californica multiple nucleopolyhedrovirus infection. Nat Commun 15 , 7711 (2024). https://doi.org/10.1038/s41467-024-52193-w
Download citation
Received : 25 January 2024
Accepted : 28 August 2024
Published : 04 September 2024
DOI : https://doi.org/10.1038/s41467-024-52193-w
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
By submitting a comment you agree to abide by our Terms and Community Guidelines . If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing: Translational Research newsletter — top stories in biotechnology, drug discovery and pharma.

IMAGES
VIDEO
COMMENTS
The significance of these viral infections to scientists is their potential impact on research, which varies depending on route of infection (i.e., natural versus experimental infection), host factors (i.e., mouse strain, age, immune competency), and virus factors (i.e., virulent versus avirulent strain; Fox et al., 2006; Percy and Barthold ...
Experimental infection of adult mice with MAdV-1 is most often subclinical and persistent but can cause fatal haemorrhagic encephalomyelitis with neurological symptoms, including tremors, seizures, ataxia and paralysis in susceptible C57BL/6 and DBA/2J mice . BALB/c mice are relatively resistant to this condition.
INTRODUCTION. Both investigative and veterinary staffs monitor the health and well-being of mice that are used in research. Indeed, this level of responsibility and care is mandated by the Public Health Service based on the Guide for the Care and Use of Laboratory Animals (National Research Council. 2011).The Guide is "intended to assist investigators in fulfilling their obligation to plan ...
Mouse infection experiments. Heterozygous K18-hACE2 C57BL/6J mice (strain 2B6.Cg-Tg(K18-ACE2)2Prlmn/J), 129 mice (strain 129S2/SvPasCrl or 129S1/SvImJ) and C57BL/6 (strain 000664) mice were ...
Intranasal infection of mice with 10 4-CFU dose of hvKp led to minor reduction in bacterial load in the airways in nociceptor-ablated mice at 24 hpi as ... mice (fig. S4B). Next, we performed the CRKP infection experiments with and without AM depletion and still observed the increased recruitment trend of Ly6C hi monocyte and neutrophils in the ...
EXPERIMENTAL MODEL. Modeling SARS-CoV-2 infection in mice. ... To characterize the susceptibility of hACE2 mice to SARS-CoV-2 infection, young (4.5-week-old) and old (30-week-old) female hACE2 and ...
a A cartoon shows schedule of the intranasal infection. The Sftpc-hACE2-N F/F mice were each intranasally infected with 1 × 10 5 TCID 50 of SARS-CoV-2 GFP/ΔN virus, and were sacrificed to ...
Mouse infection models have revealed a key protective role of the gut microbiota. Experimental mice with a complex (but specific pathogen-free, so-called "SPF") microbiome are in most cases resistant against S. Tm infection.
Mouse models have been a particularly important resource for studying infection by a wide range of human pathogens and are widely used for preclinical screening of vaccines/therapies due to their reproducibility, low cost, and ease of experimental manipulation. Although mouse models are not appropriate for studying some aspects of pathogen ...
Erwinia persicinus (E. persicina) is a plant pathogenic bacterial species that was previously isolated from a case of human infection. This study aimed to create an experimental infection protocol for E. persicina in laboratory mice. Seventy-two adult mice were divided into four groups (18 animal/group): the control group (G1), the group ...
In-bred strains of mice are commonly used to model pathogenic infections due to their cost and utility. In order to understand better the nature of experimental tuberculosis in mice, we infected BALB/c mice with a virulent field isolate of Mycobacterium bovis. Mice were sacrificed at intervals in order to visualise the pathological lesions in ...
Erwinia persicinus (E. persicina) is a plant pathogenic bacterial species that was previously isolated from a case of human infection.This study aimed to create an experimental infection protocol for E. persicina in laboratory mice. Seventy-two adult mice were divided into four groups (18 animal/group): the control group (G1), the group infected with E. persicina (G2), the group immune ...
The experimental infection with the isolated infectious LSDV could serve as a platform for future vaccine evaluation study using an LSDV challenge model. Lumpy skin disease (LSD) is one of the ...
Inflammatory and pathological changes in Escherichia coli ...
In conclusion, it is clear that the mouse genetic background influences the Leishmania infection outcome and this feature must be taken into account when designing experiments. Although BALB/c mice will not die of VL (differently from untreated humans), they can be used to study the immunopathology changes occurring during VL.
The objective of this study was to test the ability of bovine viral diarrhea virus (BVDV) to infect mice. Two mice each were either mock infected or inoculated with one of three BVDV strains by the intraperitoneal (IP) (n = 8) or intranasal (IN) (n = 8) route. All mice were euthanized at day 7 postinfection (p.i.).
(1) Background. A definition of healthcare-associated infections is essential also for the attribution of the restorative burden to healthcare facilities in case of harm and for clinical risk management strategies. Regarding M. chimaera infections, there remains several issues on the ecosystem and pathogenesis. We aim to review the scientific evidence on M. chimaera beyond cardiac surgery, and ...
Staphylococcus aureus infections of mice. When designed to recapitulate human disease, animal studies with infectious agents aim to provide experimental proof for the molecular basis of pathogenesis, the establishment of protective immunity and the molecular mechanisms whereby immunity is achieved (33-35).Over the past forty years, infectious diseases research championed the mouse as a model ...
Detection of CD4 + T cells activation via FCM and antibody levels through ELISA. (A) Immunization and infection processes in mice and the implementation of separate assays.Flow chart (B) and analysis (C) of the expression of IFN-γ + CD4 + T cells and IL-4 + CD4 + T cells in spleen.(D) The levels of sIgA in ILF and IgG in the serum of mice after the second immunization.
The global spread of monkeypox virus has raised concerns over the establishment of novel enzootic reservoirs in expanded geographic regions. We demonstrate that although deer mice are permissive to experimental infection with clade I and II monkeypox viruses, the infection is short-lived and has limited capability for active transmission.
The most common cause of ARDS is pulmonary infection, ... T. & Braun, A. Noninvasive measurement of pulmonary function in experimental mouse models of airway disease. Lung 199, 255-261.
Assistant professor of pediatrics - infectious disease Dr. Katherine King and postdoctoral fellow Dr. Katie Matatall discuss their work revealing that long-l...
Low-dose EBV infection of humanized mice results in asymptomatic, persistent infection. ... Experimental infection of NOD/SCID mice reconstituted with human CD34+ cells with Epstein-Barr virus.
The present study was undertaken to compare the parasitemia of B. microti infection in BALB/c and F1 (B10 x CBA) mice by two different methods: intraperitoneal injection of parasites or infection by the oral route. In both groups, experimental mice were inoculated with 5 x 10 (7) infected erythrocytes in 100 microliters of blood.
Methodology in experimental infections of mice with Hymenolepis nana. "natural" infection technique based on the uptake of eggs with feed is described. The ratio of the number of parasites recovered 14 days post infection to the number of eggs given (q ratio) is close to 1 if few eggs are given (= 5 per mouse).
The central and necessary role of IL-10 in protecting against severe inflammatory pathology has been clearly shown in many models of experimental infection with intracellular pathogens using Il-10 ...
Virulence assessment of six major pathogenic Candida ...
Experimental infection of mice with the microfilariae of Onchocerca lienalis Parasitology. 1982 Oct:85 (Pt 2):283-93. ... but marked differences were demonstrated between male CBA mice of different ages. After infection with 5 000 microfilariae the recovery of parasites from the ears increased rapidly to a peak at day 35 when 10% of the ...
For transduction experiments, mice were randomly grouped (n = 3 to 9 per group as indicated) and injected with baculoviruses through tail vein with a dose of 4.4 × 10 4 IU per gram body weight ...